TP53 Point Mutation, from molecular mechanisms to therapeutic strategies

The TP53 gene, also known as the p53 gene, is one of the most critical tumor suppressor genes in the human genome. Discovered in 1979 and recognized for its tumor-suppressive function in 1989, TP53 has since been regarded as the “guardian of the genome” and remains a central focus of cancer research.
When TP53 gene mutations occur, this finely tuned regulatory system collapses. In human cancers, TP53 mutations are predominantly point mutations, particularly missense mutations. These alterations can arise as:
l Somatic TP53 mutations, acquired during tumor development;
l Germline TP53 gene mutations, which are inherited and associated with Li-Fraumeni syndrome.
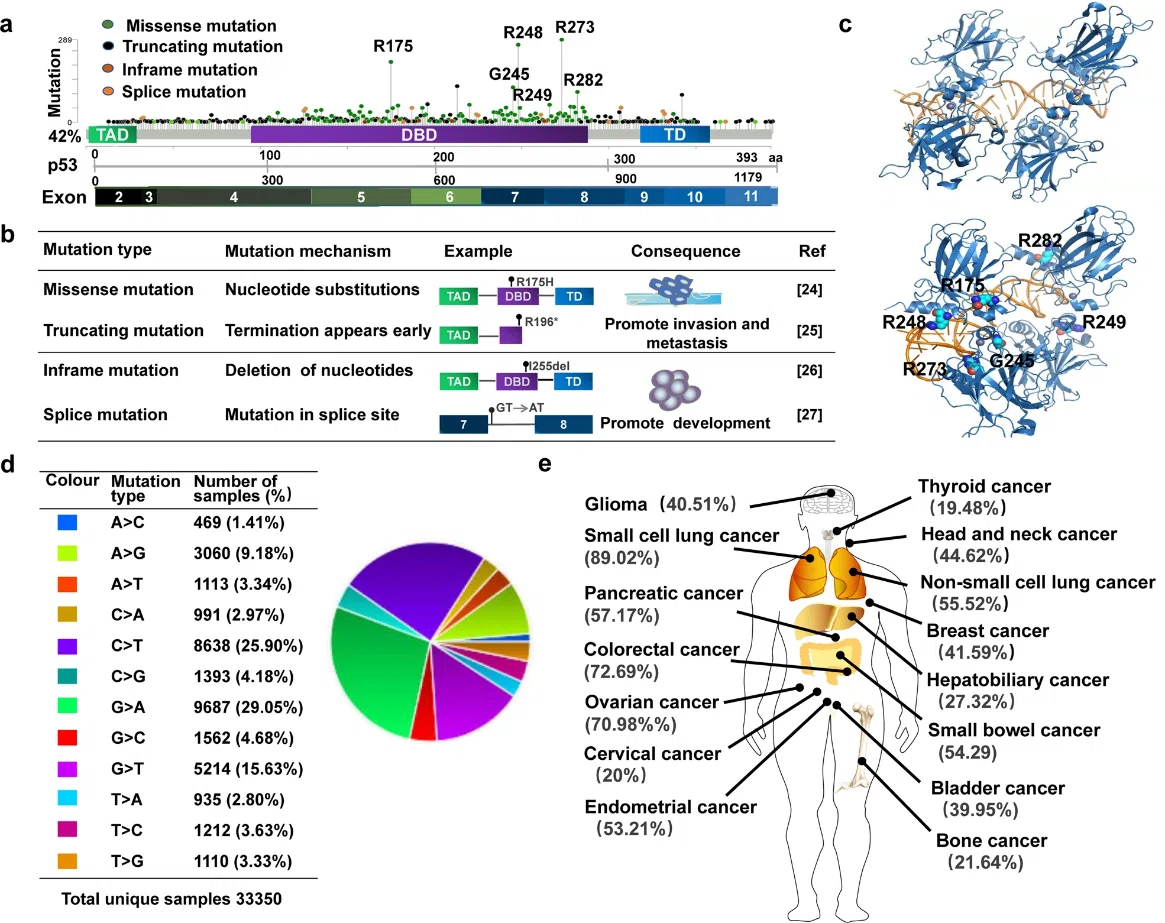
Fig. 1. General characteristics of mutp53. [1]
l Increased cancer susceptibility, with germline TP53 mutation carriers showing a lifetime cancer risk exceeding 90%;
Unlike kinase-driven oncogenic mutations, targeting TP53 mutations remains challenging due to the complexity of the p53 signaling network and the heterogeneity of TP53 gene mutations. Current research directions include:
1. Reactivation of mutant p53
Small molecules, metal ion coordination, and structural stabilizers are being developed to restore wild-type–like conformation and activity in specific TP53 mutations.
2. Synthetic lethality approaches
TP53-deficient tumors rely on alternative DNA repair pathways. By targeting these compensatory mechanisms, synthetic lethality strategies selectively eliminate cancer cells.
In addition, tumors harboring TP53 mutations often exhibit a high mutational burden, making them more responsive to immunotherapies such as immune checkpoint inhibitors.
EDITGENE has established a comprehensive portfolio of TP53 gene-edited cell lines, including TP53 point mutation and TP53 knockout models. These standardized and functionally validated cell models support mechanistic studies, drug discovery, and translational cancer research.
Point Mutation Cell Line
| Catalog | Cell Name | Gene | PM Site/Locus | Functional Domain | Order Now |
|---|
| EDC03134 | HCT116 | TP53 | p.V157F | DNA-binding domain (DBD) | Order |
| EDC03127 | HCT116 | TP53 | p.V157G | DNA-binding domain (DBD) | Order |
| EDC03143 | HCT116 | TP53 | p.R158G | DNA-binding domain (DBD) | Order |
| EDC03089 | HCT116 | TP53 | p.R158C | DNA-binding domain (DBD) | Order |
| EDC03088 | HCT116 | TP53 | p.R158H | DNA-binding domain (DBD) | Order |
| EDC03092 | HCT116 | TP53 | p.R158L | DNA-binding domain (DBD) | Order |
| EDC03063 | HCT116 | TP53 | p.H179R | DNA-binding domain (DBD) | Order |
| EDC03133 | HCT116 | TP53 | p.C242Y | DNA-binding domain (DBD) | Order |
| EDC03093 | HCT116 | TP53 | p.G245S | DNA-binding domain (DBD) | Order |
| EDC03099 | HCT116 | TP53 | p.G245C | DNA-binding domain (DBD) | Order |
| EDC03095 | HCT116 | TP53 | p.G245D | DNA-binding domain (DBD) | Order |
| EDC03132 | HCT116 | TP53 | p.G245V | DNA-binding domain (DBD) | Order |
| EDC03094 | HCT116 | TP53 | p.R248W | DNA-binding domain (DBD) | Order |
| EDC03171 | HCT116 | TP53 | p.R248L | DNA-binding domain (DBD) | Order |
| EDC03087 | HCT116 | TP53 | p.R249G | DNA-binding domain (DBD) | Order |
| EDC03086 | HCT116 | TP53 | p.R249W | DNA-binding domain (DBD) | Order |
| EDC03118 | HCT116 | TP53 | p.R249M | DNA-binding domain (DBD) | Order |
| EDC03117 | HCT116 | TP53 | p.R249S | DNA-binding domain (DBD) | Order |
| EDC03109 | HCT116 | TP53 | p.R273C | DNA-binding domain (DBD) | Order |
| EDC03170 | HCT116 | TP53 | p.R273H | DNA-binding domain (DBD) | Order |
| EDC03110 | HCT116 | TP53 | p.D281G | DNA-binding domain (DBD) | Order |
| EDC03100 | HCT116 | TP53 | p.R282G | DNA-binding domain (DBD) | Order |
| EDC03621 | HAP1 | TP53 | p.P72R | Proline-rich domain (PRD) | Order |
| EDC03619 | HAP1 | TP53 | c.673-36G>C | / | Order |
| EDC03620 | HAP1 | TP53 | c.376-91G>A | / | Order |
KO Cell Line
| Catalog | Cell Name | Gene | Order Now |
|---|
| EDC07854 | HCT116 | TP53 | Order |
| EDJ-KQ17910 | HEK293 | TP53 | Order |
| EDJ-KQ18086 | HeLa | TP53 | Order |
| EDJ-KQ18198 | A549 | TP53 | Order |
Leveraging the newly developed Bingo™ Platform , EDITGENE applies an optimized Prime Editing (PE7 system) to introduce precise point mutations with editing efficiencies exceeding 90% at the target locus.
This advanced gene-editing strategy enables the generation of high-efficiency, genetically stable point mutation cell lines, all of which are molecularly validated and suitable for basic research and drug discovery applications.
| Catalog | Cell Name | Gene | Gene ID | PM Site/Locus | Order Now |
|---|
| EDC03196 | HCT116 | KRAS | 3845 | p.G13C | Order |
| EDC03199 | HCT116 | KRAS | 3845 | p.Q61H | Order |
| EDC03203 | HCT116 | KRAS | 3845 | p.G12S | Order |
| EDC03206 | HCT116 | KRAS | 3845 | p.G12R | Order |
| EDC03208 | HCT116 | KRAS | 3845 | p.G12C | Order |
| EDC03038 | HCT116 | KRAS | 3845 | p.G12V | Order |
| EDC03030 | HCT116 | KRAS | 3845 | p.G13S | Order |
| EDC03035 | HCT116 | KRAS | 3845 | p.G13R | Order |
| EDC03211 | HCT116 | EGFR | 1956 | p.Q787R | Order |
| EDC03053 | HCT116 | GJB2 | 2706 | p.V13fs | Order |
| EDC03057 | HCT116 | GJB2 | 2706 | p.V37I | Order |
| EDC03054 | HCT116 | GJB3 | 2707 | p.C165T | Order |
| EDC03102 | HCT116 | SLC26A4 | 5172 | p.L676Q | Order |
| EDC03125 | HCT116 | SLC26A4 | 5172 | p.G197R | Order |
| EDC03106 | HCT116 | SLC26A4 | 5172 | p.T410M | Order |
| EDC03107 | HCT116 | SLC26A4 | 5172 | p.H723R | Order |
| EDC03080 | HCT116 | G6PD | 2539 | p.G131V | Order |
| EDC03076 | HCT116 | G6PD | 2539 | p.V291M | Order |
| EDC03181 | K562 | G6PD | 2539 | p.H32R | Order |
| EDC03077 | HCT116 | G6PD | 2539 | p.H32R | Order |
| EDC03108 | HCT116 | G6PD | 2539 | p.L342F | Order |
| EDC03152 | K562 | G6PD | 2539 | p.L342F | Order |
| EDC03168 | K562 | G6PD | 2539 | p.R459L | Order |
| EDC03134 | HCT116 | TP53 | 7157 | p.V157F | Order |
| EDC03127 | HCT116 | TP53 | 7157 | p.V157G | Order |
| EDC03143 | HCT116 | TP53 | 7157 | p.R158G | Order |
| EDC03092 | HCT116 | TP53 | 7157 | p.R158L | Order |
| EDC03063 | HCT116 | TP53 | 7157 | p.H179R | Order |
| EDC03133 | HCT116 | TP53 | 7157 | p.C242Y | Order |
| EDC03093 | HCT116 | TP53 | 7157 | p.G245S | Order |
| EDC03095 | HCT116 | TP53 | 7157 | p.G245D | Order |
| EDC03132 | HCT116 | TP53 | 7157 | p.G245V | Order |
| EDC03171 | HCT116 | TP53 | 7157 | p.R248L | Order |
| EDC03118 | HCT116 | TP53 | 7157 | p.R249M | Order |
| EDC03117 | HCT116 | TP53 | 7157 | p.R249S | Order |
| EDC03109 | HCT116 | TP53 | 7157 | p.R273C | Order |
| EDC03170 | HCT116 | TP53 | 7157 | p.R273H | Order |
| EDC03110 | HCT116 | TP53 | 7157 | p.D281G | Order |
| EDC03100 | HCT116 | TP53 | 7157 | p.R282G | Order |
| EDC03101 | HCT116 | AKT1 | 201 | p.E17K | Order |
| EDC03055 | HCT116 | ESR1 | 2099 | p.Y537N | Order |
| EDC03214 | HCT116 | ESR1 | 2099 | p.Y537S | Order |
| EDC03075 | HCT116 | FGFR3 | 2261 | p.R248C | Order |
| EDC03175 | HCT116 | FGFR3 | 2261 | p.Y373C | Order |
| EDC03105 | HCT116 | HRAS | 3265 | p.Q61K | Order |
| EDC03111 | HCT116 | HRAS | 3265 | p.G13R | Order |
| EDC03081 | HCT116 | HRAS | 3265 | p.Q61R | Order |
| EDC03112 | HCT116 | HRAS | 3265 | p.G12D | Order |
| EDC03179 | HCT116 | IDH1 | 3417 | p.R132H | Order |
| EDC03184 | HCT116 | IDH2 | 3418 | p.R172M | Order |
| EDC03176 | HCT116 | KIT | 3815 | p.V559A | Order |
| EDC03120 | HCT116 | KIT | 3815 | p.V560D | Order |
| EDC03119 | HCT116 | MAP2K1 | 5604 | p.P124S | Order |
| EDC03121 | HCT116 | MAP2K1 | 5604 | p.E203K | Order |
| EDC03177 | HCT116 | NRAS | 4893 | p.Q61R | Order |
| EDC03123 | HCT116 | NRAS | 4893 | p.G12S | Order |
| EDC03074 | HCT116 | NRAS | 4893 | p.G12C | Order |
| EDC03122 | HCT116 | NRAS | 4893 | p.G13R | Order |
| EDC03072 | HCT116 | NRAS | 4893 | p.G13D | Order |
| EDC03079 | HCT116 | PTEN | 5728 | p.R130G | Order |
| EDC03115 | HCT116 | PTEN | 5728 | p.R233* | Order |
| EDC03126 | HCT116 | PTEN | 5728 | p.R130L | Order |
| EDC03129 | HCT116 | PTEN | 5728 | p.R173H | Order |
| EDC03114 | HCT116 | RB1 | 5925 | p.R579* | Order |
| EDC03116 | HCT116 | RB1 | 5925 | p.R251* | Order |
| EDC03033 | HCT116 | EGFR | 1956 | p.S768I | Order |
| EDC07731 | A549 | G3BP1 | 10146 | p.S373G | Order |
| EDC03185 | A549 | USP24 | 23358 | NC_000001.11:g.55132659 T>C & 55132661 T>C | Order |
| EDC03187 | A549 | PDZD7 | 79955 | p.K790E | Order |
| EDC03189 | AC16 | GLA | 2717 | p.D155E | Order |
| EDC03141 | HEPG2 | SLC37A4 | 2542 | NC_000011.10:g.119025190C>A | Order |
| EDC03192 | HCT116 | TOPBP1 | 11073 | p.T1105A | Order |