-
CRISPR Knockout KitCRISPR Point Mutation KitKI Enhancer Drug
-
Precision Mutation Cell PanelsWild Type Cell Line
-
-
 Knockout Cell Series - Lipoprotein Stimulus
Knockout Cell Series - Lipoprotein Stimulus
Cellular response to lipoprotein particle stimulus refers to the series of intracellular responses initiated by cells upon sensing stimulation from any type of lipoprotein particle (including low-density lipoprotein LDL, high-density lipoprotein HDL, very low-density lipoprotein VLDL, and oxidized lipoprotein oxLDL).
Gene knockout models built around these processes provide powerful tools to dissect the distinct roles of lipoprotein particles across different physiological and pathological contexts.
Based on the CRISPR technology platform, EDITGENE provides a series of cell products and services covering gene knockout related to multiple types of lipoprotein particles, supporting research on lipid transport mechanisms and the exploration of related disease targets.
Lipoprotein particles are lipid–protein complexes composed of lipids and apolipoproteins, and can be classified into chylomicrons, VLDL, LDL, and HDL based on density and function. These particles transport cholesterol, triglycerides, and other lipids through the circulatory system, supporting essential biological processes such as energy supply, membrane synthesis, and signaling molecule production.
Different classes of lipoprotein particles serve distinct roles; for example, LDL primarily delivers cholesterol to peripheral tissues, whereas HDL mediates reverse cholesterol transport. The structural composition and dynamic remodeling of these particles directly influence their biological functions. Dysregulation can lead to lipid accumulation, inflammation, and metabolic imbalance, contributing to atherosclerosis and other metabolic disorders.
Studying lipoproteins at the particle level provides a more refined understanding of lipid metabolism and its role in disease.
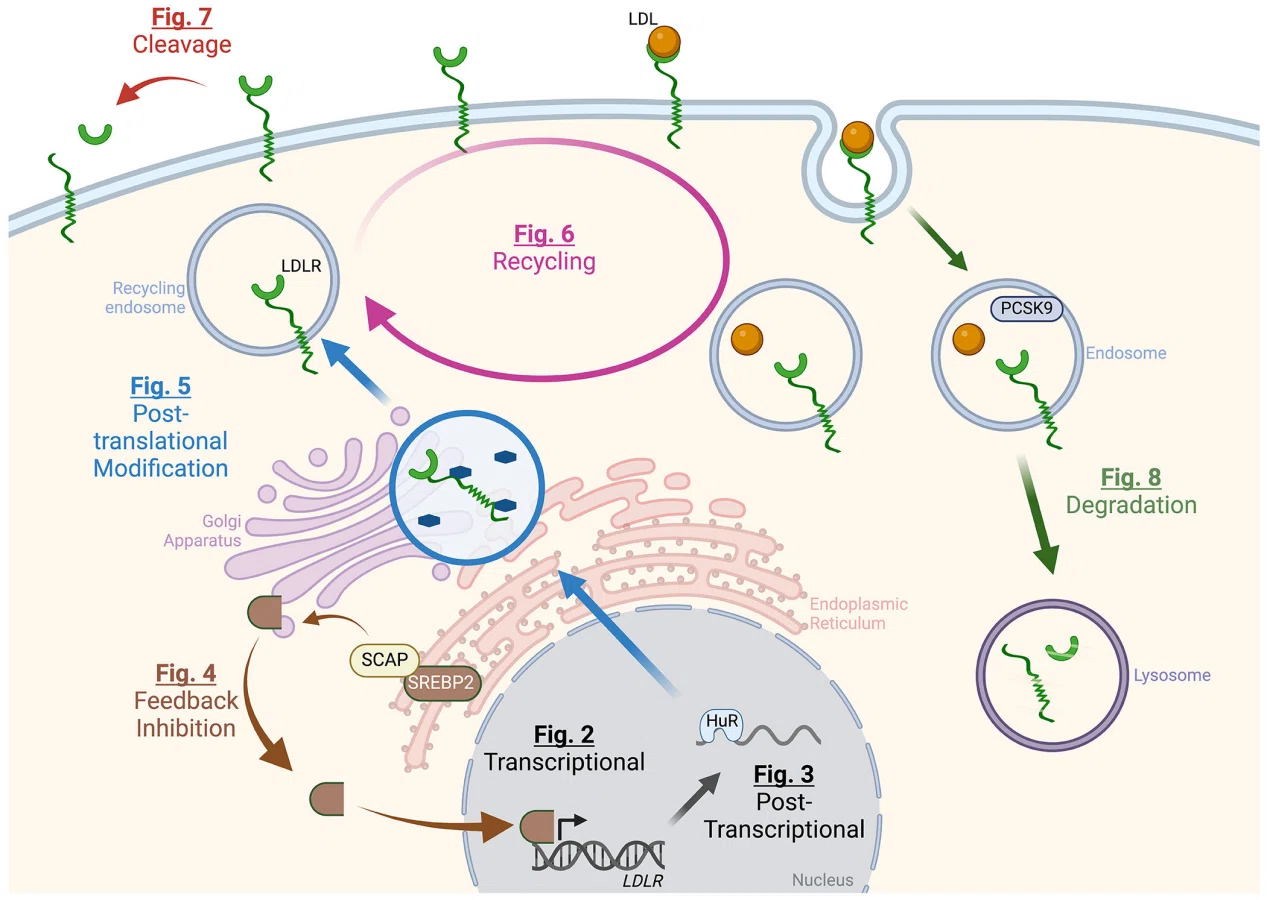
(Schmidt et al., Circ Res, 2025)
Research on Lipoprotein Particles has evolved from simple classification toward systematic analysis of particle composition, structure, and dynamic remodeling. The focus extends beyond functional differences among particle classes to include how structural changes influence function under various physiological and disease conditions. Integrative approaches enable the identification of key regulators that shape lipoprotein behavior.
In practice, this field often combines particle profiling, functional screening, and molecular perturbation strategies to systematically investigate lipoprotein-associated genes and assess their roles in disease contexts. This workflow—from structural characterization to functional impact and mechanistic insight—positions Lipoprotein Particles as a key entry point for studying lipid metabolism.
With CRISPR-based approaches, researchers can precisely manipulate genes involved in lipoprotein assembly, modification, and clearance, improving study efficiency and facilitating target validation.
1. Atherosclerosis and Cardiovascular Research
Examine structural variations of lipoprotein particles such as LDL and HDL and their roles in cholesterol deposition and plaque formation.
2. Dyslipidemia and Metabolic Disorders
Analyze how changes in particle composition and distribution contribute to hyperlipidemia, obesity, and diabetes.
3. Lipoprotein Remodeling Mechanisms
Investigate structural remodeling processes, including lipid exchange and apolipoprotein dynamics, and their functional consequences.
4. Inflammation and Immune Studies
Explore the roles of specific particles, such as oxidized LDL, in inflammatory signaling and immune activation.
5. Genetic Lipid Disorders
Study the impact of mutations in key genes (e.g., APOB, APOE) on lipoprotein particle formation and function using gene editing models.
To support Lipoprotein Particles research, EDITGENE provides knockout cell models targeting key genes involved in lipoprotein structure and function. These standardized models enable efficient investigation of particle biology and regulatory mechanisms.
The following products cover major lipoprotein-associated targets, supporting applications in lipid metabolism research and disease studies.
-
Cat.No: EDC07969
species: Human
cell_name: HAP1
gene_name: ABCA1
gene_id: 19
specs: 1×10⁶cells
-
Cat.No: EDC09412
species: Human
cell_name: HAP1
gene_name: TESK1
gene_id: 7016
specs: 1×10⁶cells
-
Cat.No: EDC07598
species: Mouse
cell_name: BV-2
gene_name: Trem2
gene_id: 83433
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ37
species: Human
cell_name: Hep-G2
gene_name: ITGB1
gene_id: 3688
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ240
species: Human
cell_name: HEK293T
gene_name: NPC1
gene_id: 4864
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ273
species: Human
cell_name: HEK293
gene_name: LDLR
gene_id: 3949
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ446
species: Human
cell_name: HEK293
gene_name: AKT1
gene_id: 207
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ528
species: Human
cell_name: HEK293
gene_name: SOCS5
gene_id: 9655
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ578
species: Human
cell_name: HEK293
gene_name: MYD88
gene_id: 4615
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ590
species: Human
cell_name: HEK293
gene_name: SYK
gene_id: 6850
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ593
species: Human
cell_name: HEK293
gene_name: TICAM1
gene_id: 148022
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ817
species: Human
cell_name: HEK293
gene_name: ITGB1
gene_id: 3688
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ864
species: Human
cell_name: HEK293
gene_name: RXRA
gene_id: 6256
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1046
species: Human
cell_name: HEK293
gene_name: CD81
gene_id: 975
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1115
species: Human
cell_name: HEK293
gene_name: PPARG
gene_id: 5468
specs: 1×10⁶cells
- 1
- 2
- ...
- 10
- 11
- Next Page »
Subscribe
You can unsubscribe from these communications at any time. For more information on how to unsubscribe, our privacy practices, and how we are committed to protecting and respecting your privacy, please review our Privacy Policy.
By clicking submit below, you consent to allow EDITGENE to store and process the personal information submitted above to provide you the content requested.