Drug Target ldentification
Empowering Drug Target Discovery with CRISPR
From Functional Genomics to Precision Therapeutic Targeting
From Correlative Studies to Causal Functional Insights
The introduction of CRISPR has shifted drug target research from passive correlation-based analysis to active, function-driven intervention. Among these approaches, CRISPR-mediated gene knockout (KO) has become a cornerstone technology for target discovery and validation.
Mechanistically, the CRISPR system uses sgRNA to guide nucleases to specific genomic loci, inducing double-strand breaks and resulting in gene inactivation.
Compared with transcriptional regulation strategies, KO eliminates gene function at the genomic level, enabling direct evaluation of phenotypic outcomes in an endogenous context—thus providing clear causal evidence linking genes to disease.
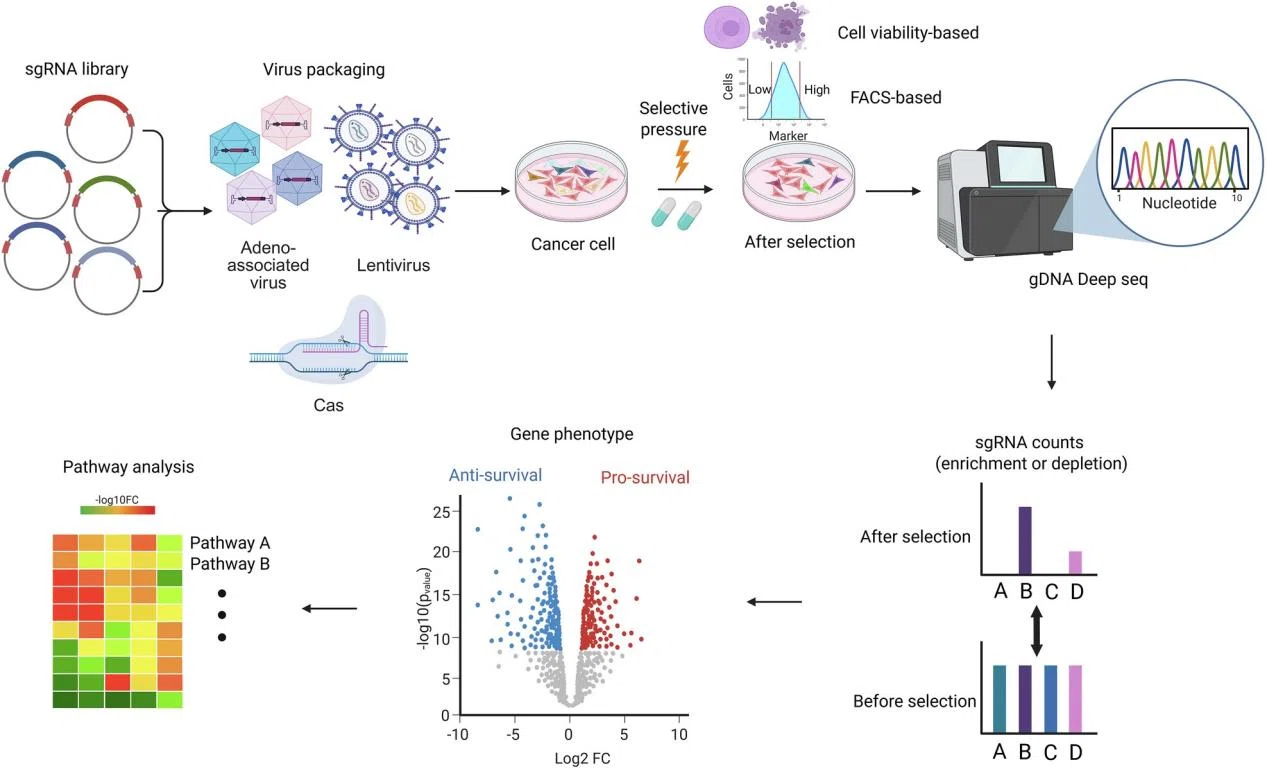
Park et al., Exp Mol Med, 2025
Furthermore, KO strategies can be expanded into high-throughput screening tools. By constructing sgRNA knockout libraries and applying selection pressures (e.g., drug treatment or phenotypic screening), researchers can systematically assess gene function at the population level. Sequencing-based analysis of sgRNA abundance enables precise identification of key functional genes.
This "gene loss → phenotypic change" framework transforms target discovery from correlation-based inference into functional validation. Particularly in complex diseases, KO models help distinguish driver genes from passenger alterations, significantly improving target identification accuracy.
From Single-Gene Perturbation to Genome-Wide Functional Interrogation
Building on the "gene loss → phenotype" framework, CRISPR screening has evolved into a multi-layered system for functional perturbation, enabling systematic gene function analysis at different scales.
CRISPR screening strategies primarily include:
These complementary approaches extend beyond simple loss-of-function analysis, enabling bidirectional regulation and comprehensive functional interrogation.
With advances in single-cell sequencing, CRISPR screening has further evolved toward higher resolution. Technologies such as Perturb-seq and CROP-seq integrate sgRNA identity with single-cell transcriptomic data, allowing researchers to dissect transcriptional changes at the single-cell level. This reveals cellular heterogeneity and dynamic regulatory processes that are not captured in bulk screening.
This integrated "perturbation–phenotype–mechanism" framework enables deeper insights into regulatory networks and mechanisms underlying disease.
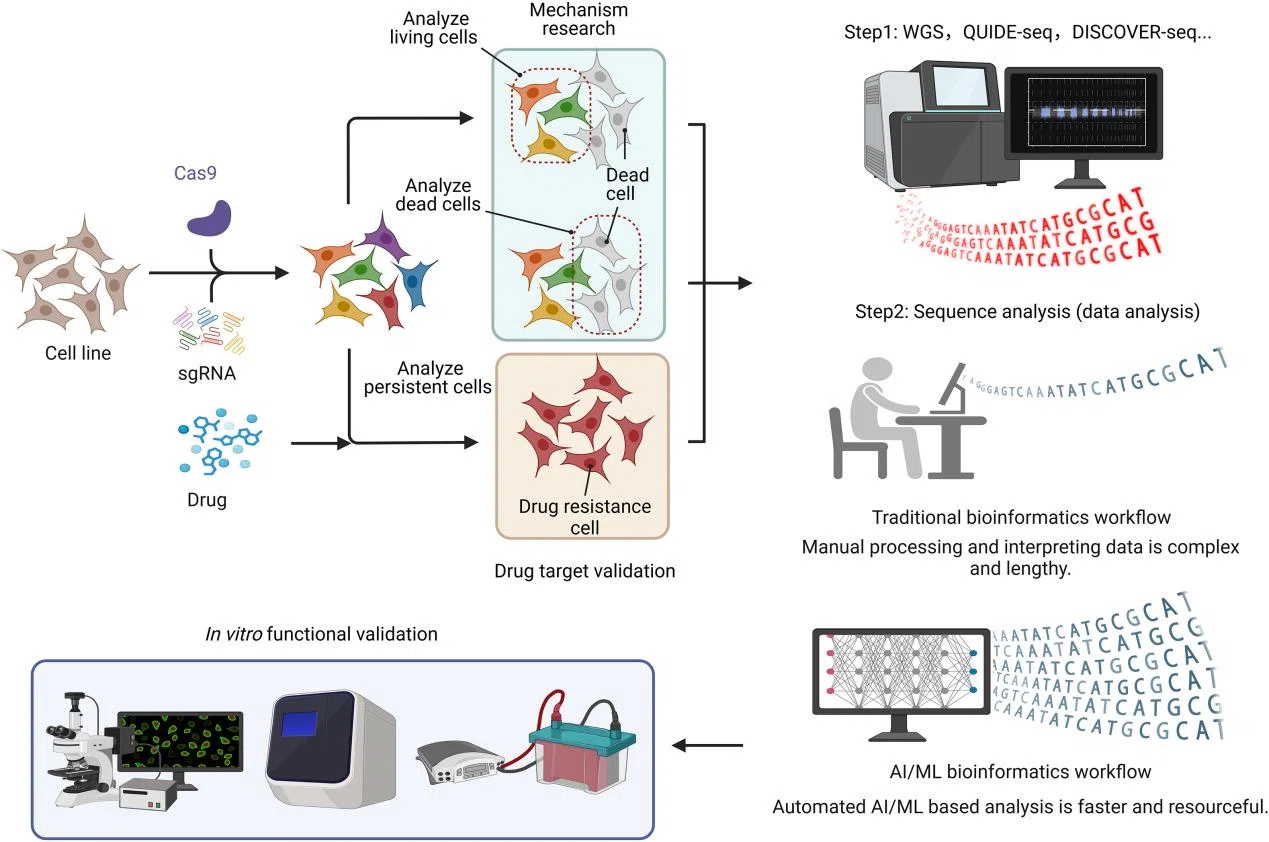
He et al., J Pharm Anal, 2025
From Candidate Genes to Mechanistic and Functional Confirmation
Accelerating the Path from Target Discovery to Drug Development
CRISPR has become deeply integrated into the entire drug development pipeline—from early target discovery and mechanistic studies to preclinical validation, biomarker identification, and patient stratification.
In oncology, genome-wide CRISPR screening has enabled the discovery of hundreds of potential therapeutic targets, including synthetic lethal genes, immune checkpoint regulators, and metabolic pathways.
In neurodegenerative, autoimmune, and infectious diseases, CRISPRa/i screening has revealed numerous gain- and loss-of-function targets. Combined with humanized models and in vivo screening, these discoveries are rapidly advancing toward clinical translation.
End-to-End Workflow for Drug Target Discovery
| Dimension | Key Scientific Question | Technology | Resources |
| Target Discovery | Which genes are associated with disease phenotypes / pathways? | CRISPR pooled screening + multi-omics integration | Genome-wide & custom CRISPR libraries |
| Functional Validation | Do candidate genes have causal roles? | CRISPR KO / CRISPRi / a perturbation | KO models & custom KO cells |
| Disease Modeling | Can mutations be recapitulated endogenously? | CRISPR knock-in (KI) | KI models & custom KI cells |
Built for Precision and Speed
Advanced Gene Editing Platforms
Integrating HES-KI and CRISPR-EDITx, our advanced platforms deliver efficient and reliable solutions for diverse editing needs.
Extensive KO Cell Line Library
Covering a wide range of genes, our extensive KO cell line library supports systematic loss-of-function studies across various disease models.
Fast Delivery for Custom Knockout Models
With a turnaround time as short as 4 weeks for custom knockout models, we significantly reduce project waiting time and accelerate your research.
Monoclonal Validation with Sequencing Confirmation
Single-clone selection combined with sequencing confirmation ensures accuracy and reproducibility for every editing outcome.
Global Project Support and Delivery
Backed by a mature global project collaboration and logistics system, we provide full-process support from technical consultation to final delivery.
Experienced Team
Expert team with over 1000 gene editing projects and experience across 300+ cell types, providing full-process support from consultation to delivery.




 Explore Cell Models
Explore Cell Models Request Project Design
Request Project Design Knockout Cell Models
Knockout Cell Models Customized Gene Knockout Services
Customized Gene Knockout Services
 HES-KI Technology Platform
HES-KI Technology Platform