-
CRISPR Knockout KitCRISPR Point Mutation KitKI Enhancer Drug
-
Precision Mutation Cell PanelsWild Type Cell Line
-
-
 Knockout Cell Series - Lipoprotein Response
Knockout Cell Series - Lipoprotein Response
Response to lipoprotein particle is a Gene Ontology term describing the biological processes through which cells or organisms respond to stimulation by lipoprotein particles, including LDL, HDL, and VLDL.
Knockout models developed around these pathways enable systematic dissection of lipid metabolism regulatory networks and help accelerate target validation.
Powered by a well-established CRISPR platform, EDITGENE offers a broad portfolio of knockout products targeting key genes involved in lipoprotein regulation, supporting applications from basic research to drug discovery.
Lipoprotein Response refers to a central biological process through which the body regulates lipid uptake, transport, and metabolism. It involves dynamic interactions among multiple classes of lipoprotein particles—such as LDL, HDL, and VLDL—as well as their associated receptors, enzymes, and signaling pathways.
By transporting hydrophobic lipids like cholesterol and triglycerides through the bloodstream, lipoproteins enable efficient energy distribution and molecular exchange between cells.
During this process, cells sense fluctuations in lipoprotein levels and activate coordinated transcriptional and signaling networks—including pathways such as SREBP, LXR, and PPAR—to maintain lipid homeostasis.
Disruption of lipoprotein metabolism can lead to cholesterol accumulation, inflammation, and systemic metabolic imbalance, contributing to the development of diseases such as atherosclerosis, obesity, diabetes, and cancer.
As a result, Lipoprotein Response represents not only a fundamental layer of metabolic regulation, but also a critical entry point for disease mechanism studies and therapeutic target discovery.
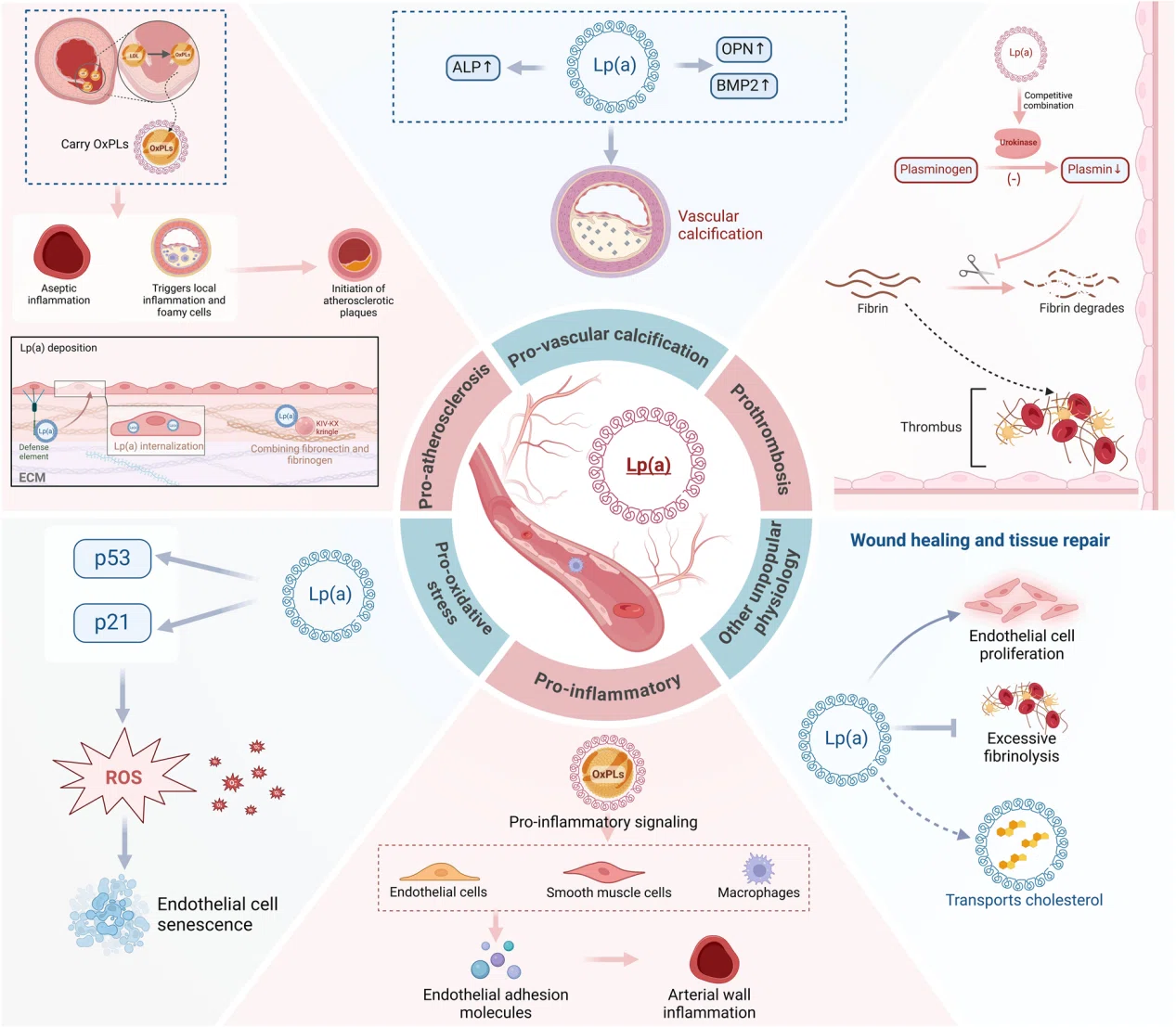
(Xu et al., Lipids Health Dis, 2025)
Research on Lipoprotein Response is shifting from single-gene or pathway-focused studies toward system-level functional analysis and regulatory network reconstruction.
Focusing on the core phenotype of abnormal lipid metabolism, research typically requires the integration of multiple regulatory factors—including receptors, transport proteins, enzymes, and transcriptional regulatory elements—to identify the critical nodes driving disease pathogenesis.
In practice, studies often combine high-throughput screening with functional validation strategies to systematically evaluate candidate genes and further characterize their context-dependent roles across different disease settings.
This workflow—from candidate discovery to functional validation and mechanistic confirmation—positions Lipoprotein Response as a critical bridge between basic research and target development.
With the advancement of CRISPR technologies, researchers can precisely perturb key regulatory factors in cell-based models, improving the efficiency and robustness of functional studies and accelerating the identification and validation of potential therapeutic targets.
1. Cardiovascular Disease Research
Investigate the roles of LDL metabolism, cholesterol accumulation, and inflammation in atherosclerosis and identify key regulatory genes.
2. Metabolic Disease Target Discovery
Explore lipid metabolism pathways, including lipoprotein uptake and reverse cholesterol transport, to uncover targets for obesity, diabetes, and dyslipidemia.
3. Inflammation and Immune Regulation
Study how lipoproteins influence inflammatory signaling, such as oxidized LDL-induced immune activation and cytokine production.
4. Cancer Metabolic Reprogramming
Examine how tumor cells adapt lipid utilization and depend on lipoprotein-mediated metabolic pathways.
5. Drug Discovery and Functional Validation
Validate lipid-related targets (e.g., LDLR, APOE, PCSK9) using gene editing and screening approaches.
To support Lipoprotein Response research, EDITGENE has developed a portfolio of knockout cell models targeting key regulators involved in lipoprotein metabolism and signaling pathways. These standardized and well-characterized models enable efficient functional studies and mechanistic exploration.
The following products cover major targets and pathway components, supporting applications in lipid metabolism research, disease modeling, and drug discovery.
-
Cat.No: EDC07796
species: Human
cell_name: HEK293
gene_name: ADAM17
gene_id: 6868
specs: 1×10⁶cells
-
Cat.No: EDC07735
species: Human
cell_name: Hep-G2
gene_name: APOE
gene_id: 348
specs: 1×10⁶cells
-
Cat.No: EDC07969
species: Human
cell_name: HAP1
gene_name: ABCA1
gene_id: 19
specs: 1×10⁶cells
-
Cat.No: EDC09412
species: Human
cell_name: HAP1
gene_name: TESK1
gene_id: 7016
specs: 1×10⁶cells
-
Cat.No: EDC07598
species: Mouse
cell_name: BV-2
gene_name: Trem2
gene_id: 83433
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ37
species: Human
cell_name: Hep-G2
gene_name: ITGB1
gene_id: 3688
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ172
species: Human
cell_name: HEK293
gene_name: APOE
gene_id: 348
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ240
species: Human
cell_name: HEK293T
gene_name: NPC1
gene_id: 4864
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ273
species: Human
cell_name: HEK293
gene_name: LDLR
gene_id: 3949
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ446
species: Human
cell_name: HEK293
gene_name: AKT1
gene_id: 207
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ528
species: Human
cell_name: HEK293
gene_name: SOCS5
gene_id: 9655
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ578
species: Human
cell_name: HEK293
gene_name: MYD88
gene_id: 4615
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ590
species: Human
cell_name: HEK293
gene_name: SYK
gene_id: 6850
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ593
species: Human
cell_name: HEK293
gene_name: TICAM1
gene_id: 148022
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ817
species: Human
cell_name: HEK293
gene_name: ITGB1
gene_id: 3688
specs: 1×10⁶cells
- 1
- 2
- ...
- 12
- 13
- Next Page »
Subscribe
You can unsubscribe from these communications at any time. For more information on how to unsubscribe, our privacy practices, and how we are committed to protecting and respecting your privacy, please review our Privacy Policy.
By clicking submit below, you consent to allow EDITGENE to store and process the personal information submitted above to provide you the content requested.