-
CRISPR Knockout KitCRISPR Point Mutation KitKI Enhancer Drug
-
Precision Mutation Cell PanelsWild Type Cell Line
-
-
 Knockout Cell Series - Mitophagy
Knockout Cell Series - Mitophagy
Mitophagy is a selective process that removes damaged mitochondria to maintain cellular homeostasis. Dysregulation of mitophagy is closely associated with neurodegenerative diseases, cardiovascular disorders, cancer, and aging-related metabolic dysfunction.
Mitophagy knockout cell lines enable precise investigation of mitochondrial quality control and disease mechanisms. Below, explore EDITGENE’s mitophagy-related KO cell models and their key research applications.
Mitophagy is a specialized form of autophagy that selectively eliminates damaged or unnecessary mitochondria. This process is essential for maintaining mitochondrial integrity, energy balance, and cellular health.
Mitophagy is mainly regulated through two major pathways:
· PINK1/Parkin-dependent pathway
Upon mitochondrial damage, PINK1 accumulates on the outer mitochondrial membrane and recruits the E3 ligase Parkin, triggering ubiquitination of mitochondrial proteins and recruitment of autophagy adaptors such as p62, NBR1, and OPTN, leading to autophagosome formation.
· Receptor-mediated pathways (PINK1/Parkin-independent)
Proteins such as BNIP3 and BNIP3L/NIX, containing LC3-interacting regions (LIRs), directly recruit autophagy machinery under conditions like hypoxia, enabling ubiquitin-independent mitophagy.
Mitophagy can be triggered by multiple conditions, including mitochondrial damage (loss of membrane potential, ROS accumulation), nutrient deprivation (AMPK activation and mTOR inhibition), hypoxia, and cellular stress.
Given its central role in mitochondrial quality control, mitophagy is critical for understanding cell survival, stress adaptation, and disease progression.
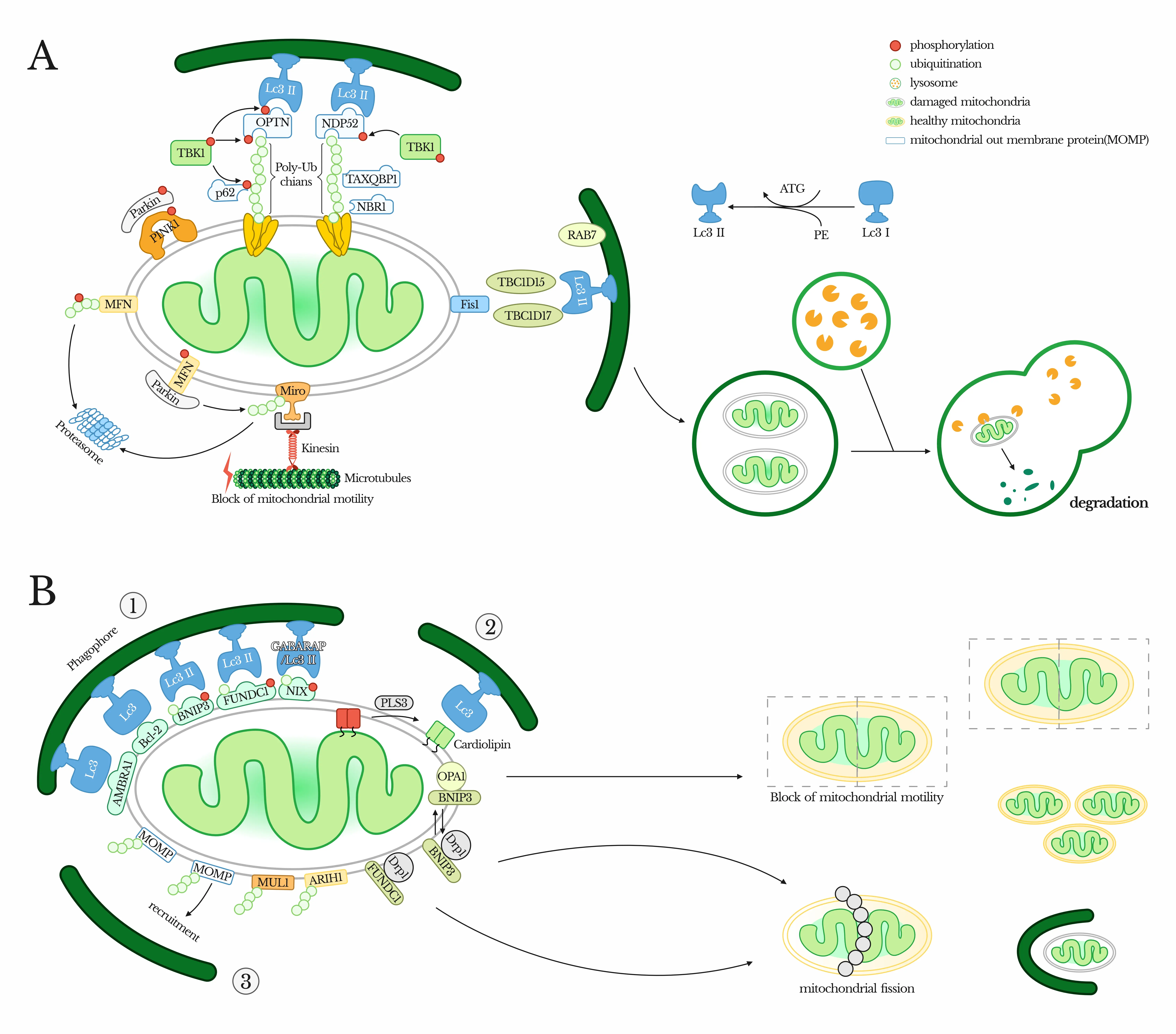
Li et al., Cell Death Dis, 2022
Mitophagy plays a key role in regulating energy homeostasis, oxidative stress response, and mitochondrial quality control, and is closely linked to multiple diseases, including cardiovascular diseases, neurodegeneration, cancer, and aging-related disorders.
Gene knockout cell models provide powerful tools to dissect mitophagy mechanisms and evaluate therapeutic strategies targeting mitochondrial dysfunction.
· Cancer & Hypoxia Models
Study hypoxia-induced mitophagy using models such as BNIP3/NIX double knockout (DKO) cells, and investigate how mitophagy regulates tumor cell survival, ferroptosis resistance, and metabolic adaptation.
· Cardiovascular Disease Models
Explore the role of mitophagy in ischemia/reperfusion injury, where it helps remove damaged mitochondria and reduce ROS accumulation, as well as its contribution to heart failure progression.
· Neurodegeneration & Aging Models
Investigate how impaired mitophagy contributes to mitochondrial dysfunction, ROS accumulation, and age-related cellular decline, providing insights into neurodegenerative diseases and aging.
· Stress & Metabolic Models
Analyze mitophagy under conditions such as nutrient deprivation, hypoxia, and toxin exposure, and study how pathways like AMPK–ULK1 signaling regulate mitochondrial turnover and cellular metabolism.
Explore the Mitophagy-Related Knockout Cell Line Collection from EDITGENE, featuring validated models targeting key regulators of mitochondrial quality control and autophagy pathways.
EDITGENE provides high-quality Mitophagy Knockout Cell Lines for studying mitochondrial dynamics, stress responses, and disease mechanisms, including models such as BNIP3/NIX DKO and PINK1/Parkin pathway genes. Both in-stock and custom gene knockout cell lines are available to support diverse mitophagy, aging, and disease research needs.
-
Cat.No: EDC08301
species: Human
cell_name: HeLa
gene_name: GBA1
gene_id: 2629
specs: 1×10⁶cells
-
Cat.No: EDC08127
species: Human
cell_name: HAP1
gene_name: TSPO
gene_id: 706
specs: 1×10⁶cells
-
Cat.No: EDC09412
species: Human
cell_name: HAP1
gene_name: TESK1
gene_id: 7016
specs: 1×10⁶cells
-
Cat.No: EDC07511
species: Mouse
cell_name: RAW 264.7
gene_name: Hk2
gene_id: 15277
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ02
species: Human
cell_name: HEK293T
gene_name: UBE2A
gene_id: 7319
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ53
species: Mouse
cell_name: MB49
gene_name: STUB1
gene_id: 56424
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ72
species: Mouse
cell_name: HT22
gene_name: PARK7
gene_id: 57320
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ306
species: Human
cell_name: HEK293
gene_name: FZD5
gene_id: 7855
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ557
species: Human
cell_name: HEK293
gene_name: CSNK2A2
gene_id: 1459
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ969
species: Human
cell_name: HEK293
gene_name: PARK7
gene_id: 11315
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1023
species: Human
cell_name: HEK293
gene_name: BNIP3
gene_id: 664
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1125
species: Human
cell_name: HEK293
gene_name: SREBF2
gene_id: 6721
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1314
species: Human
cell_name: HEK293
gene_name: DRD2
gene_id: 1813
specs: 1×10⁶cells
-
Cat.No: EDC07938
species: Human
cell_name: HEK293
gene_name: ADCY10
gene_id: 55811
specs: 1×10⁶cells
-
Cat.No: EDJ-KQ1486
species: Human
cell_name: HEK293
gene_name: NOD2
gene_id: 64127
specs: 1×10⁶cells
- 1
- 2
- ...
- 13
- 14
- Next Page »
Subscribe
You can unsubscribe from these communications at any time. For more information on how to unsubscribe, our privacy practices, and how we are committed to protecting and respecting your privacy, please review our Privacy Policy.
By clicking submit below, you consent to allow EDITGENE to store and process the personal information submitted above to provide you the content requested.