CRISPR/Cas System Principles and the Differences Between Cas9 and Cas12a

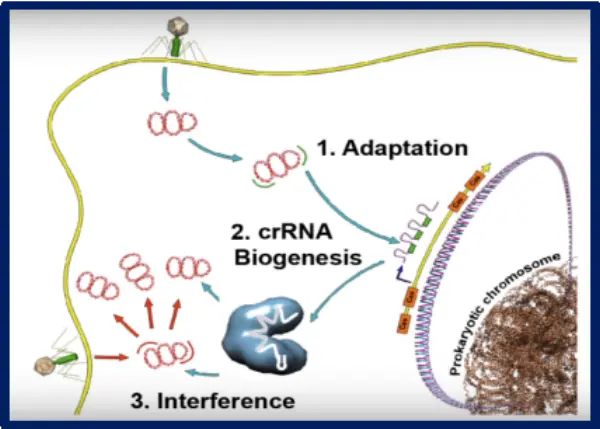
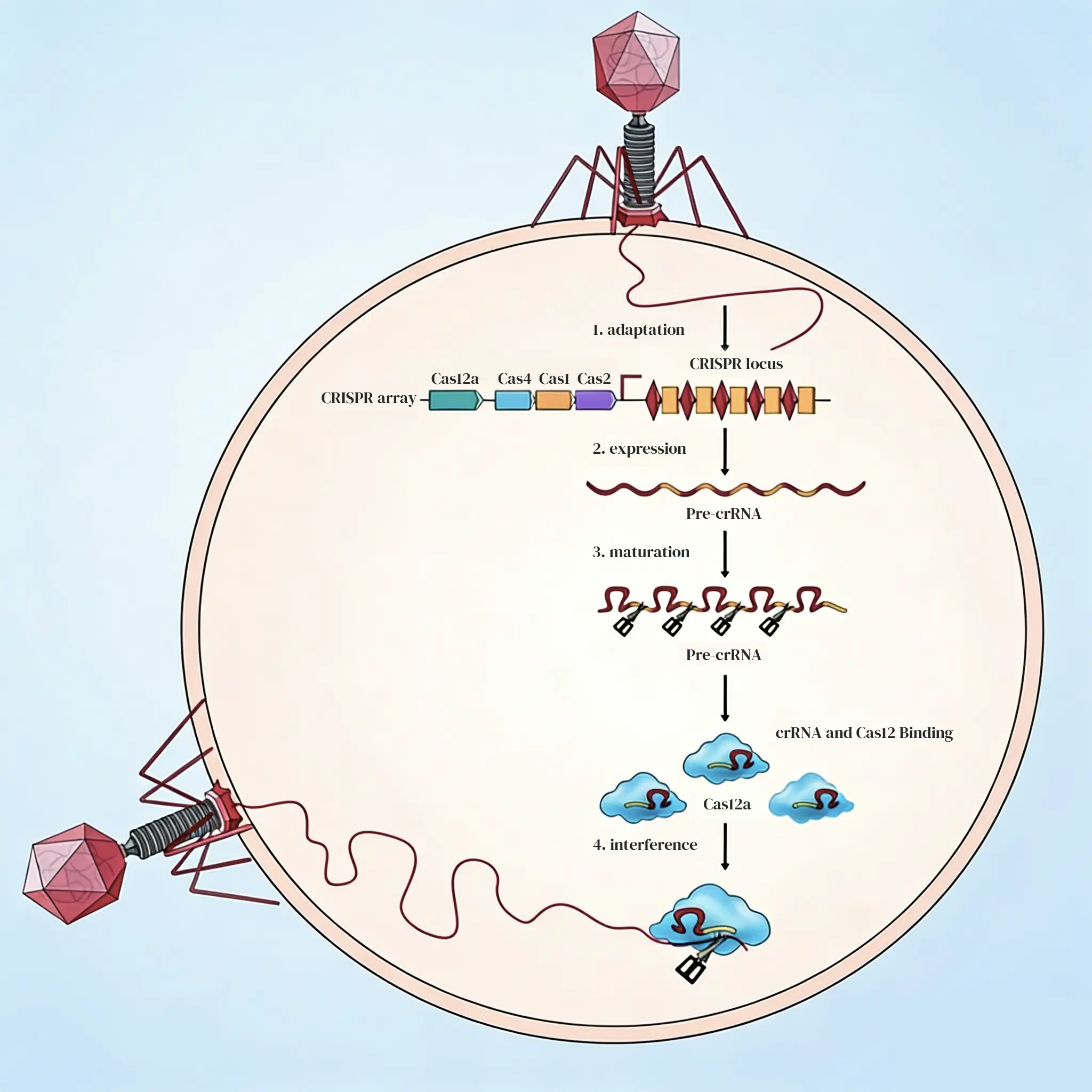
The CRISPR/Cas system is an adaptive immune system used by bacteria and archaea to defend themselves against foreign genetic material. The functionality of this system is divided into three distinct stages:
Stage 1: Adaptation
Bacteria identify PAM (Protospacer Adjacent Motif) sites within the invading nucleic acids via specific Cas proteins. The protospacer sequence is then integrated into the bacterium's own CRISPR array.
Stage 2: Expression and Maturation
The CRISPR array is transcribed into pre-crRNA (precursor RNA), which is enzymatically processed into mature crRNA. This crRNA then binds with Cas proteins to form a functional effector complex.
Stage 3: Interference
The crRNA guides the complex to recognize the PAM target on the invading nucleic acid, triggering the endonuclease activity of the Cas protein to cleave the target DNA sequence.
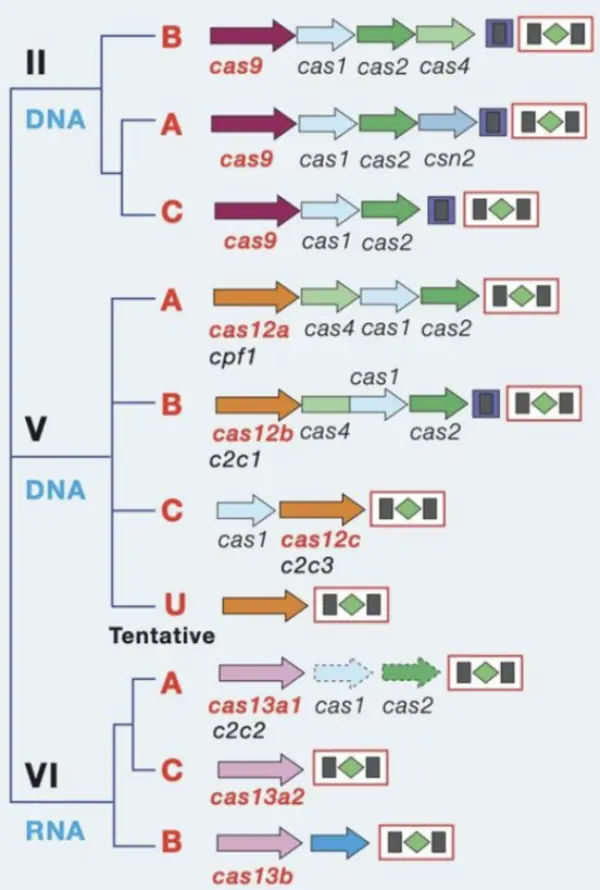
CRISPR/Cas systems are categorized into two classes and six types:
Class 1 includes Types I, III, and IV. These systems function through multi-Cas protein complexes and account for 90% of identified CRISPR/Cas loci.
Class 2 includes Types II (Cas9), V (Cas12), and VI (Cas13). These systems function via a single effector Cas protein and account for 10% of identified CRISPR/Cas loci, making them ideal tools for gene editing and in vitro diagnostics.
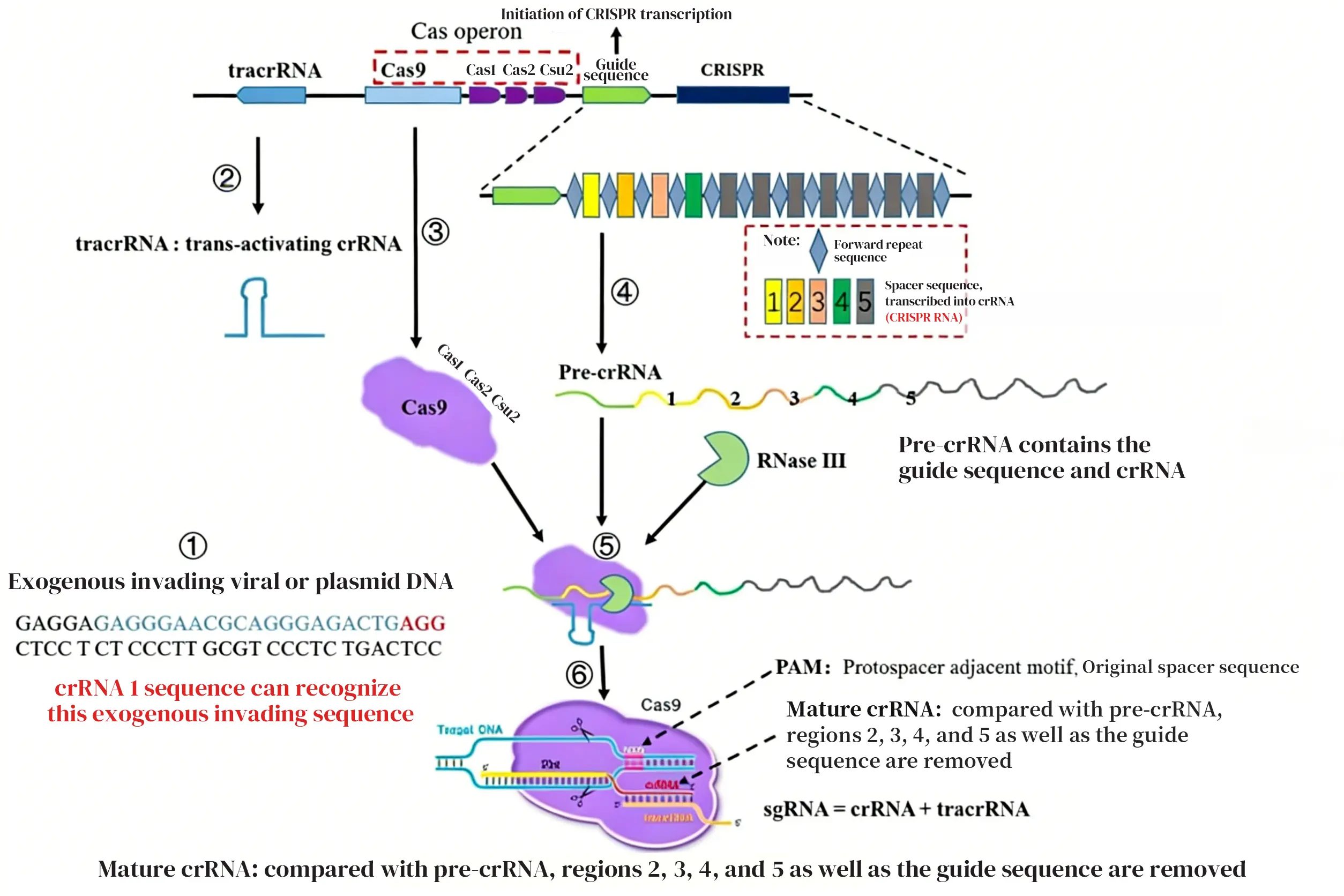
1. crRNA
In the case of Cas9, upon the invasion of exogenous nucleic acids, the bacterium transcribes tracrRNA while the Cas9 protein is simultaneously transcribed and translated. Cas1, Cas2, and Csn2 will attach to the Cas9 protein. Subsequently, the CRISPR array, regulated by the leader sequence, is transcribed into pre-crRNA.
The pre-crRNA and tracrRNA form a double-stranded RNA through complementary base pairing and assemble with the Cas9 protein into a complex. Finally, based on the type of invading nucleic acid, the corresponding spacer sequence is selected, and RNase III is utilized to cleave irrelevant sequences, resulting in the formation of a mature sgRNA (single guide RNA).
Because Cas12a possesses intrinsic ribonuclease activity, once pre-crRNA is transcribed, Cas12a can directly process pre-crRNA into its mature form without the involvement of tracrRNA or RNase III. In addition, Cas12a exhibits RNA-guided DNase activity, enabling it to cleave the target DNA.
EDITGENE provides crRNA design and synthesis services. With its proprietary crRNA design strategy, the detection limit can be improved to the attomole level, while the assay time is reduced to as short as ten minutes. This approach significantly enhances the sensitivity and cleavage activity of CRISPR-based detection.
2. PAM Sites and Domains
For Cas9, the target DNA’s PAM sequence is 5’-NGG-3’, located downstream of the spacer region on the non-target strand. Cas9’s nuclease consists of an HNH nuclease domain and a RuvC-like nuclease domain, both of which participate in DNA cleavage. The HNH domain cleaves the DNA strand complementary to the crRNA, while the RuvC-like domain cuts the non-complementary strand. The cleavage site lies between the third and fourth nucleotides upstream of the PAM within the original spacer sequence, producing a blunt-ended double-strand break (DSB).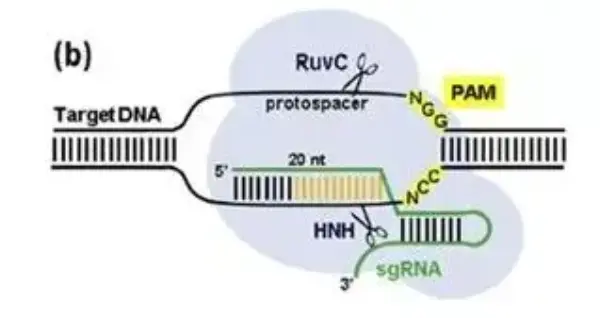
Cas12a recognizes T-rich PAM sequences, such as 5’-TTTN-3’ for LbCas12a and 5’-TTN-3’ for FnCas12a, located upstream of the spacer region on the non-target strand.
Cas12a adopts a bilobed structure composed of a REC lobe and a Nuc lobe, with nuclease activity localized in the Nuc lobe. The Nuc lobe contains the RuvC domain, the PI domain, and the WED domain. The RNase site responsible for processing its own crRNA resides in the WED domain, whereas the DNase site for DNA cleavage is located at the interface between the Nuc and RuvC domains. The RuvC domain contains the catalytic center, enabling precise recognition of the target sequence and accurate DNA cleavage under Mg²⁺ catalysis, generating staggered DSBs with sticky ends.
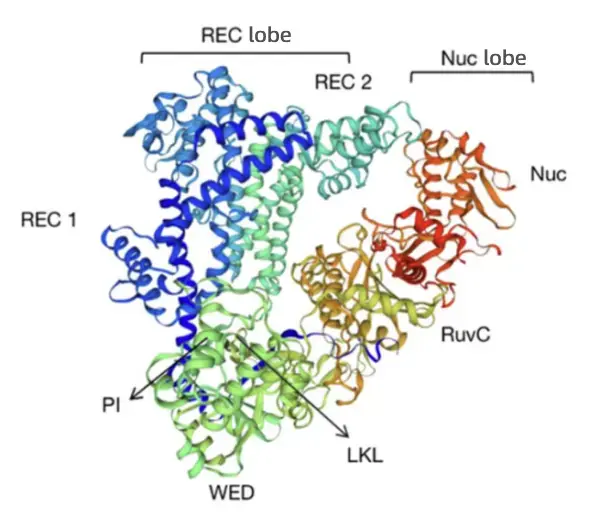
3. Mechanism of Action
When Cas9 cleaves DNA, it generates a double-strand break (DSB), triggering the cell’s DNA damage repair machinery.
Two major repair pathways exist in cells. One is non-homologous end joining (NHEJ), which can introduce small insertions, deletions, or substitutions. If the number of inserted or deleted bases is not a multiple of three, the resulting amino acid sequence produced during translation will differ completely from the original, effectively achieving gene knockout (KO).
The other pathway is homology-directed repair (HDR), which restores the DNA sequence to its pre-break state. However, NHEJ is generally the dominant repair mechanism in cells. Even when DNA is repaired via HDR, the site can still be recognized by sgRNA and subjected to repeated cleavage.
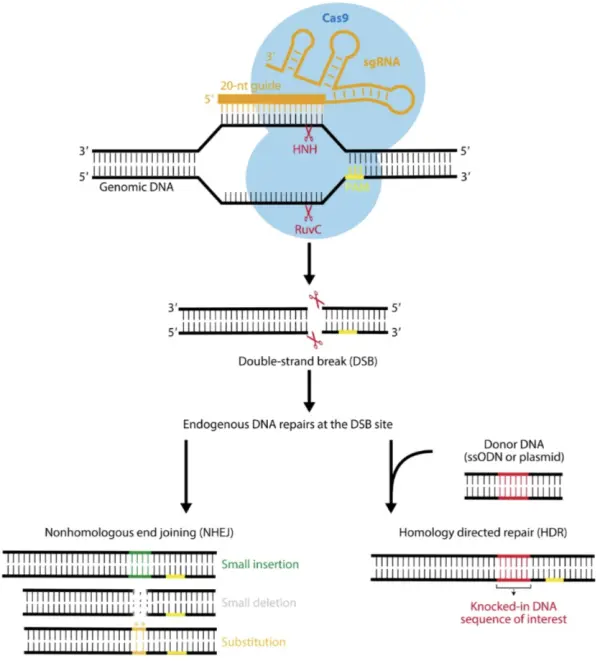
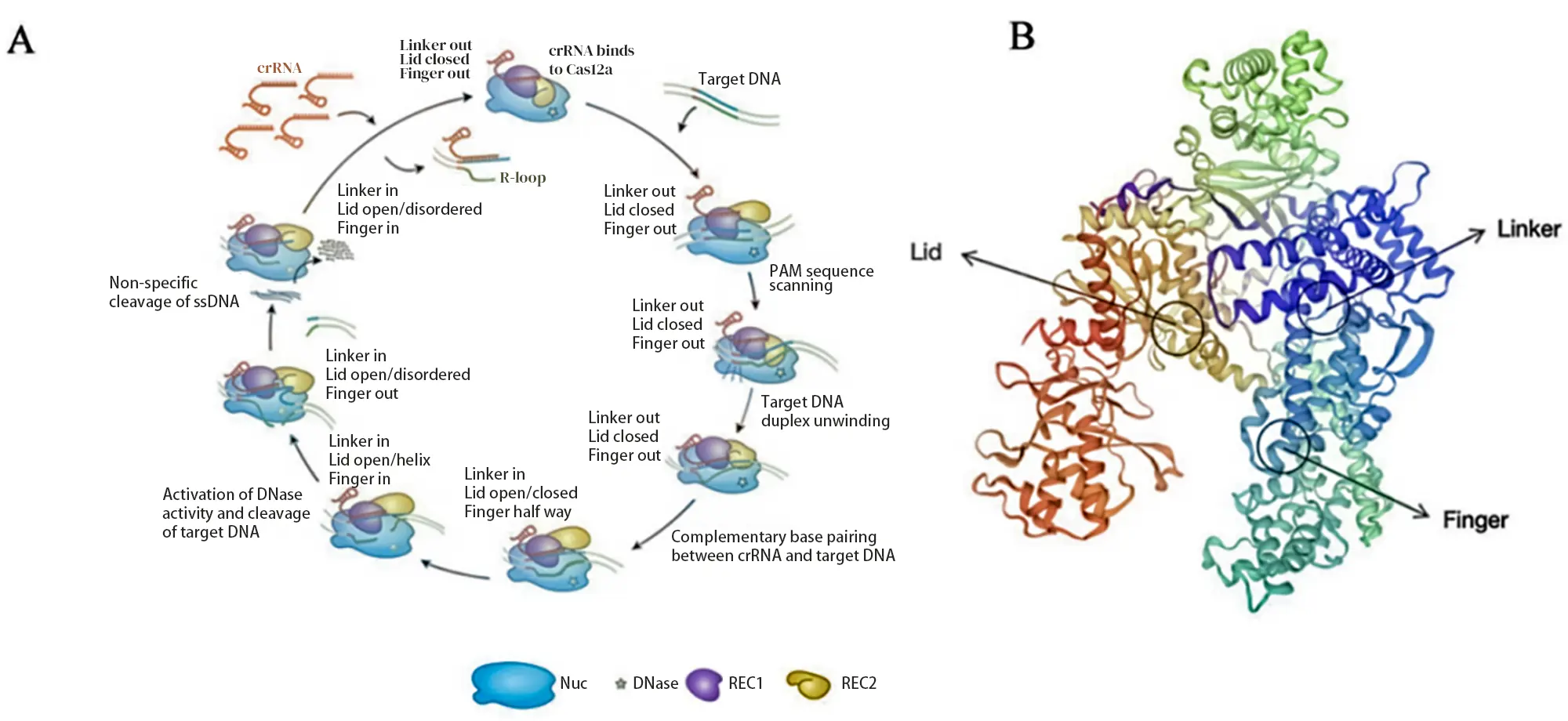
Cas12a’s RuvC domain contains several specialized structural elements:
REC Linker (Linker): a looped peptide connecting the REC1 and REC2 domains.
Lid: regulates the closure of Cas12a’s catalytic pocket.
Finger: a helical, finger-like structure within REC1.
Before crRNA hybridizes with the target DNA, the Lid remains closed, and the Linker and Finger are inactive. Upon crRNA–DNA hybridization, the Lid opens, making the Cas12a catalytic pocket accessible. After Cas12a cleaves the double-stranded DNA, the Lid undergoes a conformational change to form an α-helix, interacting with the hybridized crRNA.
The Linker and Finger move the REC domain away from the RuvC domain. At this point, the distal portion of the dsDNA substrate dissociates from the complex, exposing the catalytic site, which can then indiscriminately cleave nearby single-stranded DNA—a phenomenon called trans-cleavage.
This non-specific cleavage activity can be harnessed in detection methods: by combining it with probes, researchers can develop CRISPR-based assays that are simpler, faster, and more precise for in vitro detection.
| Feature | Cas9 | Cas12a |
|---|
| Discovery year | 2012 | 2015 |
| Cas family type | Type II | Type V |
| Protein size | 1000-1600 amino acids | ~1300 amino acids |
| gRNA | crRNA and tracrRNA | crRNA |
| Pre-crRNA processing | Requires host RNase III and tracrRNA | Processed by intrinsic RNase activity |
| PAM sequence | G-rich PAM | T-rich PAM |
| Nuclease domains | HNH and RuvC | RuvC–Nuc |
| Cleavage pattern | Blunt ends (upstream of PAM) | Sticky ends (downstream of PAM) |
| Mechanism | Relies on cellular DNA repair after cleavage | Relies on Cas12a trans-cleavage activity |
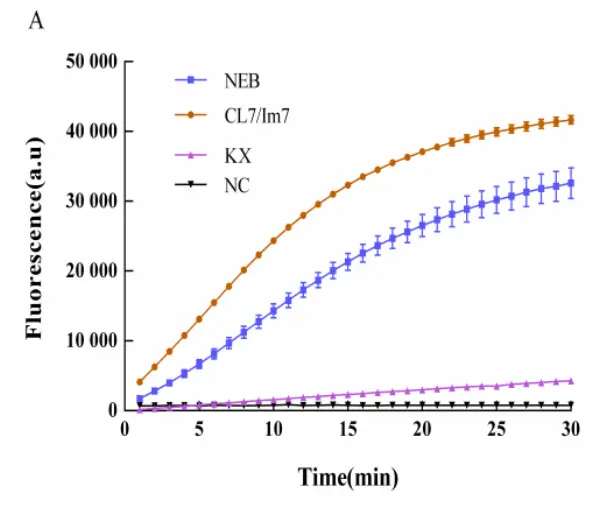
Enzyme Activity Comparison Chart
 |
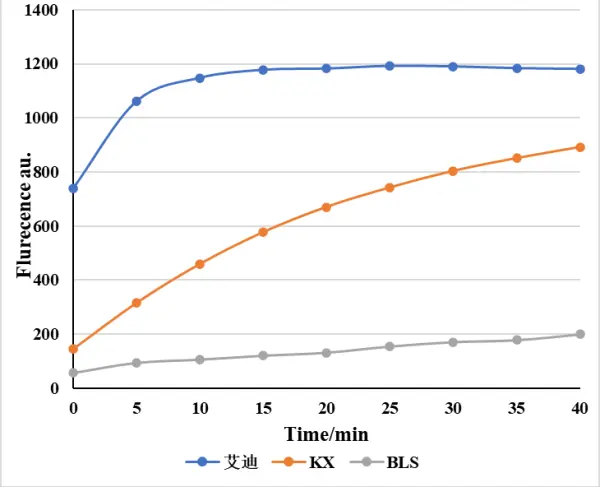 |
| LbCas12a Activity Comparison Test | LwaCas13a Activity Comparison Test |
|---|
Contact us
+ 833-226-3234 (USA Toll-free)
+1-224-345-1927 (USA)
info@editxor.com