Mitochondrial autophagy, also known as mitophagy, is a selective form of autophagy that specifically removes damaged or superfluous mitochondria from cells. It plays a central role in maintaining mitochondrial quality control, reducing the accumulation of reactive oxygen species (ROS), and regulating cellular energy metabolism and homeostasis.
In recent years, a growing body of research has demonstrated that dysregulation of mitophagy is closely associated with a wide range of diseases, including neurodegenerative disorders, cancer, cardiovascular diseases, and metabolic dysfunction. Therefore, systematically elucidating its molecular mechanisms and validating its functions using precise models has become a major focus of current research.
At its core, mitophagy involves the sequestration of specific mitochondria by autophagosomes, followed by their delivery to lysosomes for degradation. This process typically includes several key steps:
● Damage recognition: Cells detect mitochondrial dysfunction, such as loss of membrane potential or elevated ROS levels
● Signal tagging: Target mitochondria are labeled via ubiquitination or receptor proteins
● Autophagy recruitment: Autophagy-related proteins, such as LC3, are recruited to the mitochondrial surface
● Autophagosome formation and degradation: Damaged mitochondria are engulfed and ultimately degraded
Classification of Mitophagy
Based on distinct molecular mechanisms, mitophagy can be broadly classified into two categories:
1. Ubiquitin-dependent pathways (e.g., the PINK1/Parkin axis)
2. Receptor-mediated pathways (e.g., BNIP3, NIX, FUNDC1)
These pathways are activated under different stress conditions and work in concert to maintain mitochondrial dynamics and homeostasis.
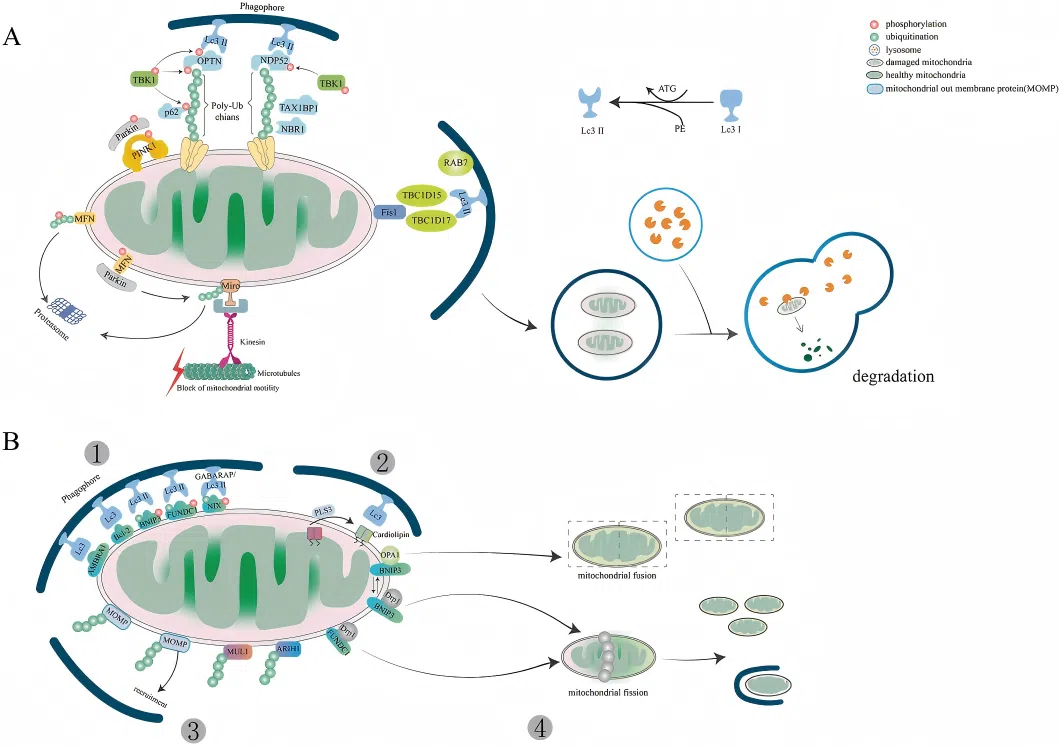
(Li et al., Cell Death Dis, 2022)
PINK1/Parkin Pathway (Figure A):
Under normal conditions, PINK1 is continuously degraded via PARL-mediated processing. Upon mitochondrial damage, however, PINK1 becomes stabilized and recruits the E3 ubiquitin ligase Parkin to initiate mitophagy. Parkin mediates polyubiquitination of mitochondrial outer membrane proteins, which in turn leads to the recruitment of autophagy adaptor proteins such as SQSTM1/p62, NBR1, and Ambra1. These adaptors bind to LC3 through their LC3-interacting regions (LIRs), facilitating the engulfment of damaged mitochondria into forming autophagosomes, which ultimately fuse with lysosomes for degradation.
Receptor-Mediated Pathway (Figure B):
BNIP3 and BNIP3L/NIX also contain LIR motifs. Their expression is induced by HIF-1α under hypoxic conditions. Unlike the ubiquitin-dependent pathway, these receptors directly recruit components of the autophagy machinery in a ubiquitin-independent manner, thereby promoting autophagosome formation in certain cell types.
Therapeutic and Application Prospects
Targeting mitophagy has emerged as a promising therapeutic strategy. By knocking out key genes involved in mitophagy, researchers can achieve the following:
1. Elucidation of key regulatory factors
Knockout of PINK1 or Parkin enables validation of their roles in damage recognition and ubiquitination, while knockout of BNIP3 or NIX facilitates investigation of receptor-mediated pathways.
2. Establishment of stable disease models
Knockout cell lines can mimic specific pathway deficiencies, providing valuable models to study disease mechanisms such as mitochondrial dysfunction or oxidative stress imbalance.
3. Dissection of complex signaling networks
Mitophagy is highly interconnected with apoptosis, autophagy, and metabolic pathways. Single- or multi-gene knockout strategies allow for effective analysis of causal relationships among these pathways.
4. Support for drug screening and target validation
Evaluating candidate compounds in knockout backgrounds helps determine whether their effects depend on specific pathways, thereby improving the precision of targeted drug development.
Based on these strategies, gene knockout cell models have become indispensable tools for investigating mitophagy mechanisms and hold broad application potential in disease research and targeted therapeutic development.
EDITGENE has developed a range of Mitophagy-related gene knockout cell models covering key pathway components such as PINK1, Parkin, BNIP3, and NIX, providing robust and efficient tools for disease mechanism studies and therapeutic strategy development.
References
[1]. Wang S, et al. (2023). The mitophagy pathway and its implications in human diseases. Signal Transduction and Targeted Therapy.
[2]. Sever T, et al. (2024). The Role of the PINK1-Parkin Pathway in Mitophagy and Neurodegenerative Diseases. Journal of Experimental and Basic Medical Sciences.
[3]. Li Y, et al. (2021). BNIP3L/NIX-mediated mitophagy: molecular mechanisms and implications for human disease. Cell Death & Disease.
[4]. Adriaenssens E, et al. (2025). Reconstitution of BNIP3/NIX-mitophagy initiation reveals hierarchical flexibility of the autophagy machinery. Nature Cell Biology.
[5]. Wang Q, et al. (2026). Mitophagy in the pathogenesis and management of disease. Cell Research.
Contact us
+ 833-226-3234 (USA Toll-free)
+1-224-345-1927 (USA)
info@editxor.com