OptiPrime for Prime Editing: How David R. Liu's AI Model Improves pegRNA Design and Gene Therapy Efficiency

On February 20, 2026, the team led by David R. Liu published their latest study, “Mechanistic machine learning enables interpretable and generalizable prediction of prime editing outcomes,” introducing OptiPrime, a mechanistically driven machine learning framework.
The OptiPrime architecture is built upon a detailed mechanistic decomposition of Prime Editing. The editing process is divided into multiple stages:
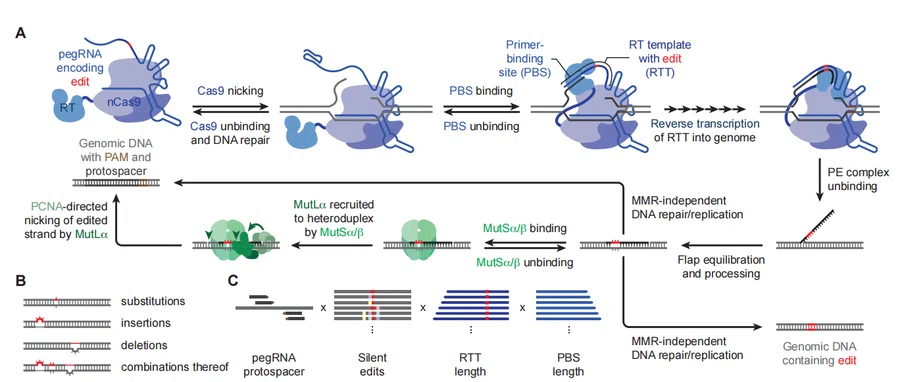
Figure 1. Design of prime editing (PE) strategies in a high-dimensional search space.
OptiPrime demonstrates strong performance, particularly in disease-relevant scenarios.
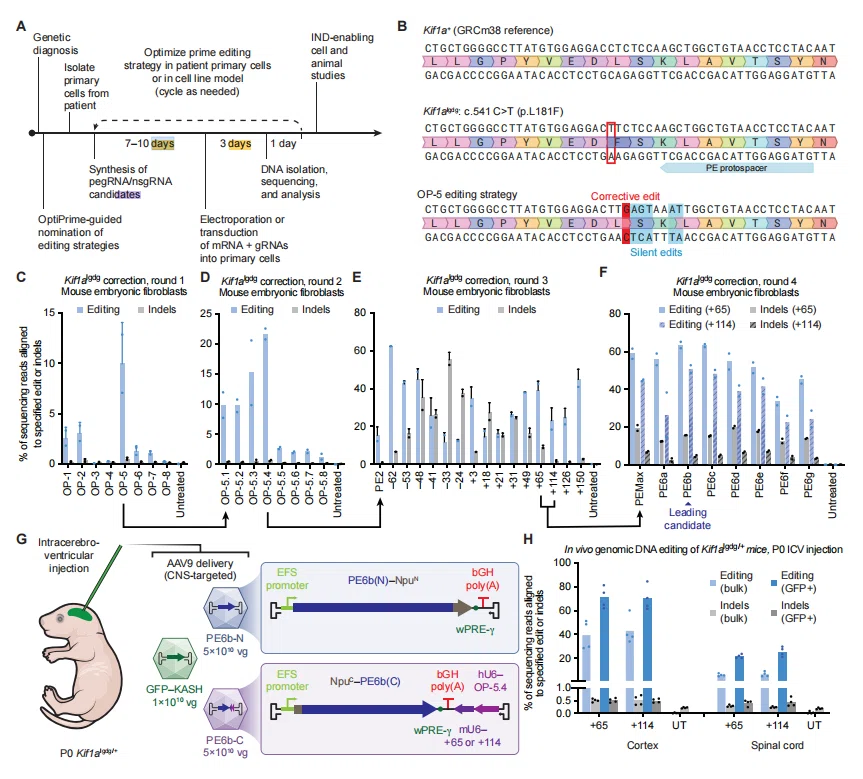
Figure 2. OptiPrime accelerates the development of corrective strategies for pathogenic mutations in vivo.
In a mouse model targeting the Kif1a “leg dragger” mutation (linked to motor dysfunction), OptiPrime nominated eight silent mutation strategies and identified OP-5 as optimal. By fine-tuning RTT and PBS combinations, only 15 pegRNAs were required to reach 22% correction efficiency in heterozygous MEFs.
From Empirical Tuning to Mechanistic Engineering
OptiPrime represents not merely an algorithmic upgrade, but a paradigm shift in pegRNA design logic.
Original link: https://www.biorxiv.org/content/10.64898/2026.02.20.706353v1
Contact us
+ 833-226-3234 (USA Toll-free)
+1-224-345-1927 (USA)
info@editxor.com