[Quality Share] sgRNA Design and Screening
![]()
The design of sgRNAs requires a comprehensive evaluation of multiple factors — including the location of candidate editing sites, GC content, and potential off-target risks — to select one or more sgRNAs with high cleavage efficiency and specificity for experimental validation.
The following principles can be used as guidelines for sgRNA design :
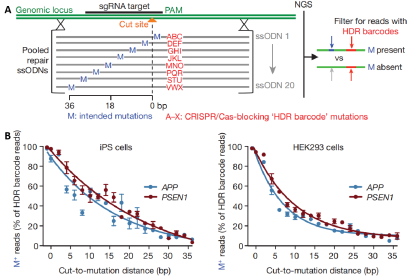
We recommend initially screening 2–4 candidate sgRNAs in parallel to identify the most efficient one for subsequet experiments.
To minimize off-target risks, the following strategies can be applied:
① Optimize sgRNA Design:
This is the most fundamental step. As the cornerstone of specificity, sgRNAs should be designed using professional software to ensure high genome-wide specific, optimal GC content and avoiding consecutive repetitive sequences.
② Use High-Fidelity Cas9 Variants
Several high-fidelity versions of Cas9 (e.g., SpCas9-HF1, eSpCas9) have been developed. These variants require more precise sgRNA-DNA pairing, significantly reducing mismatch tolerance and effectively mitigating off-target effects.
③ Improve Delivery Methods and Control Expression Duration
Transient delivery methods, such as direct introduction of ribonucleoprotein complexes (RNPs), rather than using plasmids for sustained expression, can shorten the exposure time of Cas9/sgRNA in cells and lower the chance of unintended cleavage.
④ Apply Comprehensive Off-Target Detection Techniques
Technologies like GUIDE-seq and OGM provide genome-wide off-target assessment, offering a realistic profile of gene editing safety.