How Does the CRISPR System Distinguish Self from Foreign DNA?

When conducting CRISPR-related research, have you ever wondered about this: PAM sequences consist of only a few nucleotides—for example, Cas9 recognizes 5’-NGG-3’. Such short motifs are far from rare, and similar sequences are abundant within the bacterial genome itself. If so, why doesn’t the CRISPR system mistakenly target its own genome and excise protospacers from it?
The article entitled CRISPR adaptation biases explain preference for acquisition of foreign DNA may offer valuable insights into this question.

To examine whether Cas1 and Cas2 are directly involved in spacer acquisition, the researchers designed the following experiment:
Two biological replicates were set up with 0.2% arabinose induction, two replicates without arabinose, and two corresponding empty-vector control replicates for each condition. After 16 hours of culture, the CRISPR arrays were sequenced to determine whether new spacers had been incorporated.
The results indicated that in the arabinose-induced group, 36.92% of CRISPR arrays had acquired new spacers, whereas only 2.61% showed new spacer acquisition in the non-induced group. No new spacers were detected in any of the empty-vector controls. These findings indicate that expression of Cas1 and Cas2 markedly enhances the generation of new spacers. The small number of new spacers observed in the absence of arabinose induction is likely attributable to leaky transcription of the Cas1-Cas2 system.
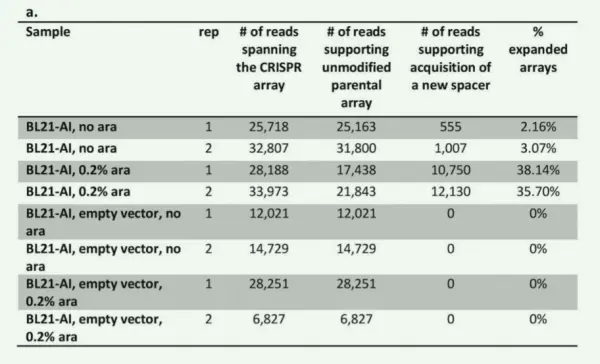
Figure 1a. Experimental validation of the association between Cas1/Cas2 and spacer acquisition
To determine the origin of newly acquired spacers, the researchers performed high-throughput sequencing based on the experimental setup described above. They found that in the group induced with 0.2% arabinose, 22.86% of the newly incorporated spacers were derived from the host chromosome, whereas only 1.8% originated from the chromosome in the non-induced group.
These data indicate that the majority of new spacers are acquired from exogenous plasmid DNA, with only a minority derived from the bacterial chromosome. Moreover, when Cas1 and Cas2 are not expressed or are expressed at low levels, the CRISPR system appears to display greater specificity toward foreign DNA during spacer acquisition.
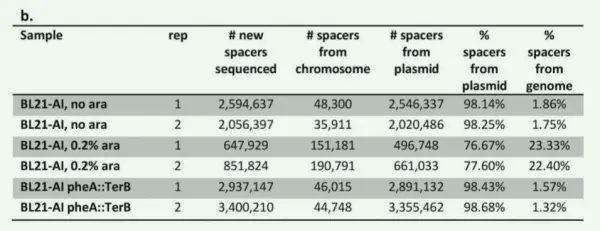
Figure 1b. Analysis of the origin of newly acquired spacers
To further explore how Cas1 and Cas2 acquire spacers from bacterial DNA, the researchers performed extensive sequencing of newly integrated spacers to examine their distribution patterns across the chromosome.
As shown in Figure 2b, PAM sequences are evenly distributed along the chromosome, indicating that PAM motifs are widespread. In contrast, the density of protospacers is clearly uneven. Figure 2a reveals that protospacer density is highest in three regions: the chromosomal origin of replication (oriC), the replication terminus (Ter), and the CRISPR locus itself, with the enrichment at the Ter region being particularly pronounced.
The Ter region marks where the two opposing replication forks converge. TerA and TerC function as unidirectional fork arrest sites, halting the replication fork that arrives first until the fork progressing from the opposite direction catches up. In Escherichia coli, the clockwise replichore (from oriC to TerA) is longer than the counterclockwise replichore (from oriC to TerC). As a result, replication forks are more likely to pause naturally at TerC than at TerA, creating a TerC/TerA bias.
Moreover, protospacer hotspots near Ter sites are asymmetric relative to the direction of replication fork movement. As illustrated in Figure 2c, strong protospacer enrichment is observed upstream of each replication fork arrest site, whereas downstream regions show comparatively lower protospacer density.
The underlying reason for this pattern will be addressed later in the study, so we will set that question aside for now. Finally, in Figure 2d, the authors inserted a native TerB site into the chromosomal pheA locus, generating a new local protospacer hotspot. This provides compelling evidence that protospacer acquisition is closely associated with replication fork stalling.
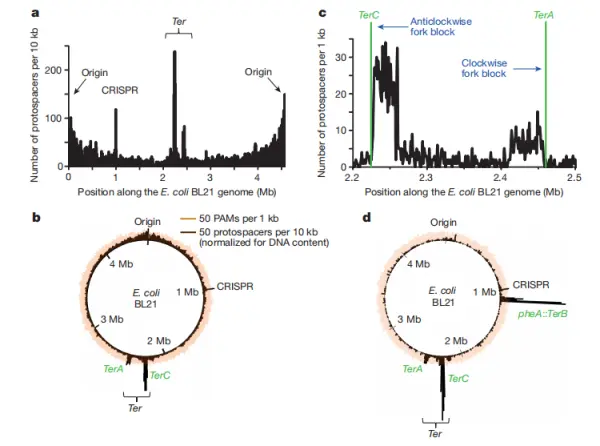
Figure 2. Chromosome-scale patterns of spacer acquisition
In Figure 3a, a pronounced protospacer hotspot is observed upstream of CRISPR I in Escherichia coli BL21-AI, whereas no such enrichment is detected upstream of CRISPR II. This is because BL21-AI lacks the leader sequence required for active adaptation and therefore cannot efficiently initiate spacer acquisition on its own; instead, integration depends on Cas1 and Cas2 supplied by pCas1+2. In contrast, as shown in Figure 3b, both CRISPR loci are active in wild-type E. coli K-12.
Based on earlier findings, the authors propose that dense protospacer regions near CRISPR arrays may cause replication fork stalling, similar to what occurs at TerA and TerC sites. This replication-associated effect may partly explain the higher protospacer density observed at TerC.
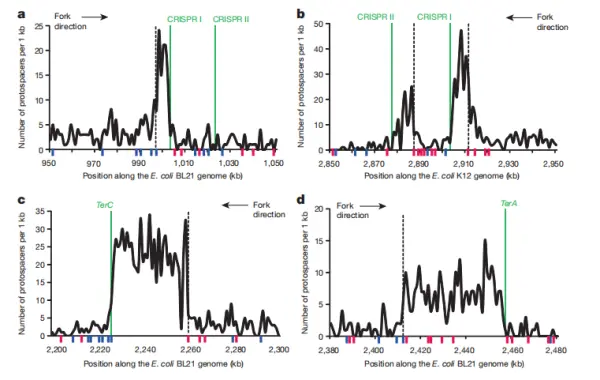
Figure 3. Spacer acquisition hotspots in E. coli BL21-AI and K-12
The authors hypothesized that DNA replication may facilitate the adaptation phase of the CRISPR system.
To test this, they performed two additional experiments based on the setup in Figure 1a. In the first group, nalidixic acid was added to inhibit DNA replication by targeting the subunits of DNA gyrase and topoisomerase IV. In the second group, the RNA polymerase inhibitor rifampicin was used to block transcription (without affecting T7 RNA polymerase-driven expression of Cas1 and Cas2).
The results showed that inhibition of DNA replication almost completely abolished the acquisition of new spacers. In contrast, blocking transcription led to a moderate reduction in new spacer incorporation, but levels remained relatively high. These findings demonstrate that spacer acquisition primarily occurs during DNA replication.
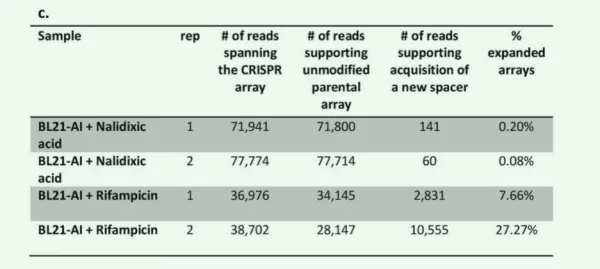
Figure 4. Experimental validation that DNA replication promotes CRISPR adaptation (Experiment 1)
To further test their hypothesis, the authors conducted an experiment using E. coli K-12 cells carrying a temperature-sensitive dnaC2 allele alongside wild-type cells. The dnaC2 mutation allows normal DNA replication at 30°C but blocks replication at 39°C.
The results showed that at 30°C, both strains exhibited detectable levels of newly acquired spacers. At 39°C, however, the dnaC2 strain showed almost no new spacer acquisition, while the wild-type K-12 cells continued to acquire new spacers. These findings provide additional evidence that DNA replication is a key driver of CRISPR adaptation.
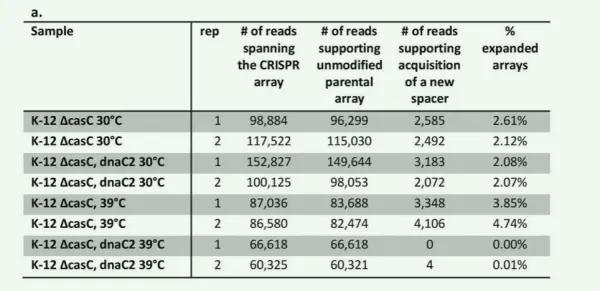
Figure 5. Experimental validation that DNA replication promotes CRISPR adaptation
To investigate whether spacer acquisition preferences are related to the position of replication forks, the authors shifted cultures of temperature-sensitive dnaC2 cells to 39°C for 70 minutes. At this temperature, DNA replication restart is blocked, and the cells contain no active replication forks.
Cas1 and Cas2 expression was then induced for 30 minutes, after which the cultures were returned to 30°C to allow synchronized initiation of DNA replication. Under these conditions, a full replication cycle in dnaC2 cells takes approximately 60 minutes. Newly acquired spacers were sequenced at 20, 40, 60, 90, and 120 minutes after replication restart.
The results showed that as the replication cycle progressed, the proportion of spacers derived from the Ter region gradually increased, peaking around 60 minutes before cycling again with the next replication round. This experiment demonstrates that during DNA replication, the Cas1-Cas2 complex preferentially acquires spacers from regions near stalled replication forks.
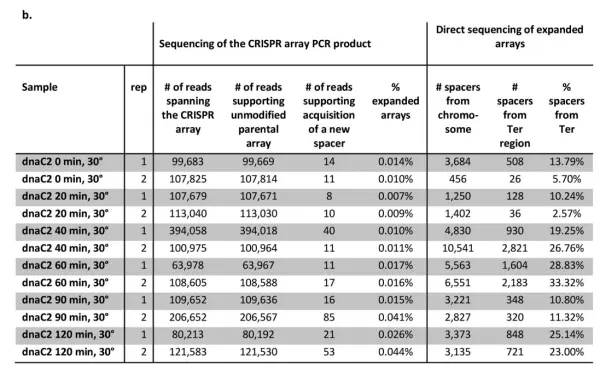
Figure 6. Experiment linking spacer acquisition preferences to replication fork positions
At this point, the pattern becomes clearer: foreign plasmids replicate frequently, generating numerous replication forks. During DNA replication, double-strand breaks are more likely to occur, making these regions prime targets for spacer acquisition by the CRISPR system. In contrast, the bacterial chromosome normally has very few exposed double-stranded DNA ends, so the likelihood of self-DNA being captured is much lower. However, when double-strand breaks do occur during chromosomal replication, there is still some chance that these regions can be incorporated as spacers.
Returning to the earlier question: why is there strong protospacer enrichment upstream of replication fork arrest sites, while downstream regions show relatively low density?
The authors observed that downstream of each fork arrest site, protospacer regions are consistently flanked by octameric GCTGGTGG motifs — the canonical Chi sequence.
Before explaining the role of Chi sites, it is important to understand the RecBCD pathway in E. coli. RecBCD functions as an additional bacterial immune mechanism and is also part of the cell’s homologous recombination–based DNA repair system. When a double-strand break occurs, RecBCD binds to the exposed DNA end, unwinds and degrades DNA until it encounters a Chi site. Upon recognizing a Chi site, RecBCD halts degradation and switches to DNA repair mode.
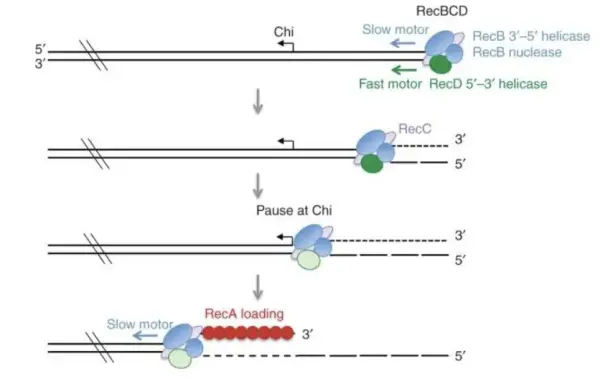
Figure 7. The RecBCD mechanism
Exogenous phage DNA is generally linear and has exposed double-stranded ends, making it highly susceptible to degradation by RecBCD. During this process, the Cas1-Cas2 complex can capture protospacers from the degrading DNA. Similarly, foreign plasmids, due to their high replication frequency, generate numerous replication forks, and DNA replication often produces double-strand breaks, making these regions readily targeted by the CRISPR system.
In bacterial chromosomes, DNA breaks are more likely to occur at stalled replication forks. During the processing of these double-strand breaks, Cas1 and Cas2 acquire spacers from RecBCD degradation intermediates. Once RecBCD encounters a Chi site, degradation stops, preventing further spacer acquisition. This explains why strong protospacer enrichment is observed upstream of replication fork arrest sites, while downstream regions show relatively low density.
Moreover, Chi sites are abundant and widely distributed across the bacterial genome, which greatly reduces the likelihood that the CRISPR system will acquire spacers from self-DNA, making foreign phages or plasmids far more likely sources for new spacers.
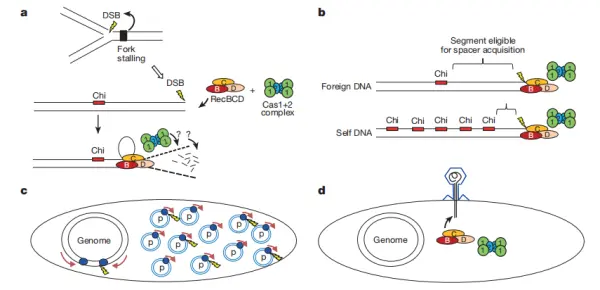
Figure 8. Mechanism of RecBCD in spacer acquisition
To further test whether spacer acquisition depends on the activity of the RecBCD complex, the authors cultured E. coli strains individually lacking recB, recC, or recD for 16 hours and then sequenced their CRISPR arrays.
The results showed that all three deletion strains exhibited a reduced overall rate of spacer acquisition. Compared with wild-type cells, the proportion of spacers derived from the host chromosome increased in the recB, recC, and recD mutants. These findings indicate that RecBCD activity plays an important role in regulating spacer acquisition.
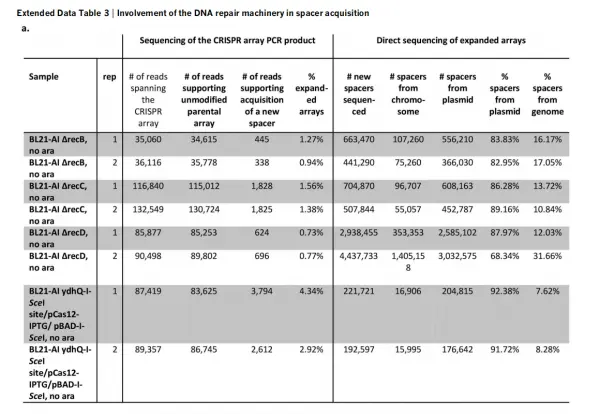
Figure 9. Correlation between spacer acquisition and RecBCD complex activity
In summary, the authors demonstrated that Cas1/Cas2–mediated acquisition of protospacers relies on DNA double-strand breaks and subsequent degradation. The Cas1-Cas2 complex captures protospacers from the intermediates generated during RecBCD-mediated DNA degradation. In bacterial chromosomes, the dense distribution of Chi sites limits the number of degradation intermediates produced by RecBCD, resulting in far fewer opportunities for spacer acquisition from self-DNA compared with exogenous plasmids or phages. Consequently, most Cas1/Cas2–acquired spacers are derived from foreign plasmids or phage DNA.
Contact us
+ 833-226-3234 (USA Toll-free)
+1-224-345-1927 (USA)
info@editxor.com